- Write my thesis
- Thesis writers
- Buy thesis papers
- Bachelor thesis
- Master's thesis
- Thesis editing services
- Thesis proofreading services
- Buy a thesis online
- Write my dissertation
- Dissertation proposal help
- Pay for dissertation
- Custom dissertation
- Dissertation help online
- Buy dissertation online
- Cheap dissertation
- Dissertation editing services
- Write my research paper
- Buy research paper online
- Pay for research paper
- Research paper help
- Order research paper
- Custom research paper
- Cheap research paper
- Research papers for sale
- Thesis subjects
- How It Works

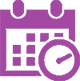
122 The Best Genetics Research Topics For Projects

The study of genetics takes place across different levels of the education system in academic facilities all around the world. It is an academic discipline that seeks to explain the mechanism of heredity and genes in living organisms. First discovered back in the 1850s, the study of genetics has come a pretty long way, and it plays such an immense role in our everyday lives. Therefore, when you are assigned a genetics research paper, you should pick a topic that is not only interesting to you but one that you understand well.
Choosing Research Topics in Genetics
Even for the most knowledgeable person in the room, choosing a genetics topic for research papers can be, at times, a hectic experience. So we put together a list of some of the most exciting top in genetics to make the endeavor easier for you. However, note, while all the topics we’ve listed below will enable you to write a unique genetic project, remember what you choose can make or break your paper. So again, select a topic that you are both interested and knowledgeable on, and that has plenty of research materials to use. Without further ado, check out the topics below.
Interesting Genetics Topics for your Next Research Paper
- Genes and DNA: write a beginners’ guide to genetics and its applications
- Factors that contribute or/and cause genetic mutations
- Genetics and obesity, what do you need to know?
- Describe RNA information
- Is there a possibility of the genetic code being confidential?
- Are there any living cells present in the gene?
- Cancer and genetics
- Describe the role of genetics in the fight against Alzheimer’s disease
- What is the gene
- Is there a link between genetics and Parkinson’s disease? Explain your answer.
- Replacement of genes and artificial chromosomes
- Explain genetic grounds for obesity
- Development and disease; how can genetics dissect the developing process
- Analyzing gene expression – RNA
- Gene interaction; eye development
- Advances and developments in nanotechnology to enable therapeutic methods for the treatment of HIV and AIDS.
- Isolating and identifying the cancer treatment activity of special organic metal compounds.
- Analyzing the characteristics in certain human genes that can withstand heavy metals.
- A detailed analysis of genotypes that is both sensitive and able to endure heavy metals.
- Isolating special growth-inducing bacteria that can assist crops during heavy metal damage and identifying lipid directing molecules for escalating heavy metal endurance in plants.
Hot and Controversial Topics in Genetics
- Is there a link between genetics and homosexuality? Explain your answer
- Is it ethical and morally upright to grow human organs
- Can DNA changes beat aging
- The history and development of human cloning science
- How addictive substances alter our genes
- Are genetically modified foods safe for human and animal consumption?
- Is depression a genetically based condition?
- Genetic diagnosis of the fetus
- Genetic analysis of the DNA structure
- What impact does cloning have on future generations?
- What is the link between genetics and autism?
- Can artificial insemination have any sort of genetic impact on a person?
- The advancements in genetic research and the bioethics that come with them.
- Is human organ farming a possibility today?
- Can genetics allow us to design and build a human to our specifications?
- Is it ethical to try and tamper with human genetics in any way?
Molecular Genetics Topics
- Molecular techniques: How to analyze DNA(including genomes), RNA as well as proteins
- Stem cells describe their potential and shortcomings
- Describe molecular and genome evolution
- Describe DNA as the agent of heredity
- Explain the power of targeted mutagenesis
- Bacteria as a genetic system
- Explain how genetic factors increase cancer susceptibility
- Outline and describe recent advances in molecular cancer genetics
- Does our DNA sequencing have space for more?
- Terminal illness and DNA.
- Does our DNA determine our body structure?
- What more can we possibly discover about DNA?
Genetic Engineering Topics
- Define gene editing, and outline key gene-editing technologies, explaining their impact on genetic engineering
- The essential role the human microbiome plays in preventing diseases
- The principles of genetic engineering
- Project on different types of cloning
- What is whole genome sequencing
- Explain existing studies on DNA-modified organisms
- How cloning can impact medicine
- Does our genetics hold the key to disease prevention?
- Can our genetics make us resistant to certain bacteria and viruses?
- Why our genetics plays a role in chronic degenerative diseases.
- Is it possible to create an organism in a controlled environment with genetic engineering?
- Would cloning lead to new advancements in genetic research?
- Is there a possibility to enhance human DNA?
- Why do we share DNA with so many other animals on the planet?
- Is our DNA still evolving or have reached our biological limit?
- Can human DNA be manipulated on a molecular or atomic level?
- Do we know everything there is to know about our DNA, or is there more?
Controversial Human Genetic Topics
- Who owns the rights to the human genome
- Is it legal for parents to order genetically perfect children
- is genetic testing necessary
- What is your stand on artificial insemination vs. ordinary pregnancy
- Do biotech companies have the right to patent human genes
- Define the scope of the accuracy of genetic testing
- Perks of human genetic engineering
- Write about gene replacement and its relationship to artificial chromosomes.
- Analyzing DNA and cloning
- DNA isolation and nanotechnology methods to achieve it.
- Genotyping of African citizens.
- Greatly mutating Y-STRs and the isolated study of their genetic variation.
- The analytical finding of indels and their genetic diversity.
DNA Research Paper Topics
The role and research of DNA are so impactful today that it has a significant effect on our daily lives today. From health care to medication and ethics, over the last few decades, our knowledge of DNA has experienced a lot of growth. A lot has been discovered from the research of DNA and genetics.
Therefore, writing a good research paper on DNA is quite the task today. Choosing the right topic can make things a lot easier and interesting for writing your paper. Also, make sure that you have reliable resources before you begin with your paper.
- Can we possibly identify and extract dinosaur DNA?
- Is the possibility of cloning just around the corner?
- Is there a connection between the way we behave and our genetic sequence?
- DNA research and the environment we live in.
- Does our DNA sequencing have something to do with our allergies?
- The connection between hereditary diseases and our DNA.
- The new perspectives and complications that DNA can give us.
- Is DNA the reason all don’t have similar looks?
- How complex human DNA is.
- Is there any sort of connection between our DNA and cancer susceptibility and resistance?
- What components of our DNA affect our decision-making and personality?
- Is it possible to create DNA from scratch under the right conditions?
- Why is carbon such a big factor in DNA composition?
- Why is RNA something to consider in viral research and its impact on human DNA?
- Can we detect defects in a person’s DNA before they are born?
Genetics Topics For Presentation
The subject of genetics can be quite broad and complex. However, choosing a topic that you are familiar with and is unique can be beneficial to your presentation. Genetics plays an important part in biology and has an effect on everyone, from our personal lives to our professional careers.
Below are some topics you can use to set up a great genetics presentation. It helps to pick a topic that you find engaging and have a good understanding of. This helps by making your presentation clear and concise.
- Can we create an artificial gene that’s made up of synthetic chromosomes?
- Is cloning the next step in genetic research and engineering?
- The complexity and significance of genetic mutation.
- The unlimited potential and advantages of human genetics.
- What can the analysis of an individual’s DNA tell us about their genetics?
- Is it necessary to conduct any form of genetic testing?
- Is it ethical to possibly own a patent to patent genes?
- How accurate are the results of a genetics test?
- Can hereditary conditions be isolated and eliminated with genetic research?
- Can genetically modified food have an impact on our genetics?
- Can genetics have a role to play in an individual’s sexuality?
- The advantages of further genetic research.
- The pros and cons of genetic engineering.
- The genetic impact of terminal and neurological diseases.
Biotechnology Topics For Research Papers
As we all know, the combination of biology and technology is a great subject. Biotechnology still offers many opportunities for eager minds to make innovations. Biotechnology has a significant role in the development of modern technology.
Below you can find some interesting topics to use in your next biotechnology research paper. Make sure that your sources are reliable and engage both you and the reader.
- Settlements that promote sustainable energy technology maintenance.
- Producing ethanol through molasses emission treatment.
- Evapotranspiration and its different processes.
- Circular biotechnology and its widespread framework.
- Understanding the genes responsible for flora response to harsh conditions.
- Molecule signaling in plants responding to dehydration and increased sodium.
- The genetic improvement of plant capabilities in major crop yielding.
- Pharmacogenomics on cancer treatment medication.
- Pharmacogenomics on hypertension treating medication.
- The uses of nanotechnology in genotyping.
- How we can quickly detect and identify food-connected pathogens using molecular-based technology.
- The impact of processing technology both new and traditional on bacteria cultures linked to Aspalathus linearis.
- A detailed analysis of adequate and renewable sorghum sources for bioethanol manufacturing in South Africa.
- A detailed analysis of cancer treatment agents represented as special quinone compounds.
- Understanding the targeted administering of embelin to cancerous cells.
Tips for Writing an Interesting Genetics Research Paper
All the genetics research topics above are excellent, and if utilized well, could help you come up with a killer research paper. However, a good genetics research paper goes beyond the topic. Therefore, besides choosing a topic, you are most interested in, and one with sufficient research materials ensure you
Fully Understand the Research Paper Format
You may write on the most interesting genetics topics and have a well-thought-out set of ideas, but if your work is not arranged in an engaging and readable manner, your professor is likely to dismiss it, without looking at what you’ve written. That is the last thing you need as a person seeking to score excellent grades. Therefore, before you even put pen to paper, understand what research format is required.
Keep in mind that part of understanding the paper’s format is knowing what words to use and not to use. You can contact our trustful masters to get qualified assistance.
Research Thoroughly and Create an Outline
Whichever genetics research paper topics you decide to go with, the key to having excellent results is appropriately researching it. Therefore, embark on a journey to understand your genetics research paper topic by thoroughly studying it using resources from your school’s library and the internet.
Ensure you create an outline so that you can note all the useful genetic project ideas down. A research paper outline will help ensure that you don’t forget even one important point. It also enables you to organize your thoughts. That way, writing them down in the actual genetics research paper becomes smooth sailing. In other words, a genetics project outline is more like a sketch of the paper.
Other than the outline, it pays to have an excellent research strategy. In other words, instead of looking for information on any random source you come across, it would be wise to have a step-by-step process of looking for the research information.
For instance, you could start by reading your notes to see what they have to say about the topic you’ve chosen. Next, visit your school’s library, go through any books related to your genetics research paper topic to see whether the information on your notes is correct and for additional information on the topic. Note, you can visit the library either physically or via your school’s website. Lastly, browse educational sites such as Google Scholar, for additional information. This way, you’ll start your work with a bunch of excellent genetics project ideas, and at the same time, you’ll have enjoyed every step of the research process.
Get Down to Work
Now turn the genetics project ideas on your outline into a genetics research paper full of useful and factual information.
There is no denying writing a genetics research paper is one of the hardest parts of your studies. But with the above genetics topics and writing tips to guide you, it should be a tad easier. Good luck!
Leave a Reply Cancel reply
- How It Works
- PhD thesis writing
- Master thesis writing
- Bachelor thesis writing
- Dissertation writing service
- Dissertation abstract writing
- Thesis proposal writing
- Thesis editing service
- Thesis proofreading service
- Thesis formatting service
- Coursework writing service
- Research paper writing service
- Architecture thesis writing
- Computer science thesis writing
- Engineering thesis writing
- History thesis writing
- MBA thesis writing
- Nursing dissertation writing
- Psychology dissertation writing
- Sociology thesis writing
- Statistics dissertation writing
- Buy dissertation online
- Write my dissertation
- Cheap thesis
- Cheap dissertation
- Custom dissertation
- Dissertation help
- Pay for thesis
- Pay for dissertation
- Senior thesis
- Write my thesis
119 Genetics Research Topics You Must Know About

Put simply, Genetics is the study of genes and hereditary traits in living organisms. Knowledge in this field has gone up over time, and this is proportional to the amount of research.
Right from the DNA structure discovery, a lot more has come out into the open. There are so many genetics research topics to choose from because of the wide scope of research done in recent years.
Genetics is so dear to us since it helps us understand our genes and hereditary traits. In this guide, you will get to understand this subject more and get several topic suggestions that you can consider when looking for interesting genetics topics.
Writing a paper on genetics is quite intriguing nowadays. Remember that because there are so many topics in genetics, choosing the right one is crucial. It will help you cut down on research time and the technicality of selecting content for the topic. Thus, it would matter a lot if you confirmed whether or not the topic you’re choosing has relevant sources in plenty.
What Is Genetics?
Before we even go deeper into genetics topics for research papers, it is essential to have a basic understanding of what the subject entails.
Genetics is a branch of Biology to start with. It is mainly focused on the study of genetic variation, hereditary traits, and genes.
Genetics has relations with several other subjects, including biotechnology, medicine, and agriculture. In Genetics, we study how genes act on the cell and how they’re transmitted from a parent to the offspring. In modern Genetics, the emphasis is more on DNA, which is the chemical substance found in genes. Remember that Genetics cut across animals, insects, and plants – basically any living organism there is.
Tips On How To Write A Decent Research Paper On Genetics
When planning to choose genetics topics, you should also make time and learn how to research. After all, this is the only way you can gather the information that will help you come up with the content for the paper. Here are some tips that can bail you out whenever you feel stuck:
Choosing the topic, nonetheless, is not an easy thing for many students. There are just so many options present, and often, you get spoilt for choice. But note that this is an integral stage/process that you have to complete. Do proper research on the topic and choose the kind of information that you’d like to apply.
Choose a topic that has enough sources academically. Also, choosing interesting topics in genetics is a flex that can help you during the writing process.
On the web, there’s a myriad of information that often can become deceiving. Amateurs try their luck to put together several pieces of information in a bid to try and convince you that they are the authority on the subject. Many students become gullible to such tricks and end up writing poorly in Genetics.
Resist the temptation to look for an easy way of gaining sources/information. You have to take your time and dig up information from credible resources. Otherwise, you’ll look like a clown in front of your professor with laughable Genetics content.
Also, it is quite important that you check when your sources were updated or published. It is preferred and advised that you use recent sources that have gone under satisfactory research and assessment.
Also, add a few words to each on what you’re planning to discuss.Now, here are some of the top genetics paper topics that can provide ideas on what to write about.
Good Ideas For Genetics Topics
Here are some brilliant ideas that you can use as research paper topics in the Genetics field:
- Is the knowledge of Genetics ahead of replication and research?
- What would superman’s genetics be like?
- DNA molecules and 3D printing – How does it work?
- How come people living in mountainous regions can withstand high altitudes?
- How to cross genes in distinct animals.
- Does gene-crossing really help to improve breeds or animals?
- The human body’s biggest intriguing genetic contradictions
- Are we still far away from achieving clones?
- How close are we to fully cloning human beings?
- Can genetics really help scientists to secure various treatments?
- Gene’s regulation – more details on how they can be regulated.
- Genetic engineering and its functioning.
- What are some of the most fascinating facts in the field of Genetics?
- Can you decipher genetic code?
- Cancer vaccines and whether or not they really work.
- Revealing the genetic pathways that control how proteins are made in a bacterial cell.
- How food affects the human body’s response to and connection with certain plants’ and animals’ DNA.
Hot Topics In Genetics
In this list are some of the topics that raise a lot of attention and interest from the masses. Choose the one that you’d be interested in:
- The question of death: Why do men die before women?
- Has human DNA changed since the evolution process?
- How much can DNA really change?
- How much percentage of genes from the father goes to the child?
- Does the mother have a higher percentage of genes transferred to the child?
- Is every person unique in terms of their genes?
- How does genetics make some of us alike?
- Is there a relationship between diets and genetics?
- Does human DNA resemble any other animal’s DNA?
- Sleep and how long you will live on earth: Are they really related?
- Does genetics or a healthy lifestyle dictate how long you’ll live?
- Is genetics the secret to long life on earth?
- How much does genetics affect your life’s quality?
- The question on ageing: Does genetics have a role to play?
- Can one push away certain diseases just by passing a genetic test?
- Is mental illness continuous through genes?
- The relationship between Parkinson’s, Alzheimer’s and the DNA.
Molecular Genetics Topics
Here is a list of topics to help you get a better understanding of Molecular genetics:
- Mutation of genes and constancy.
- What can we learn more about viruses, bacteria, and multicellular organisms?
- A study on molecular genetics: What does it involve?
- The changing of genetics in bacteria.
- What is the elucidation of the chemical nature of a gene?
- Prokaryotes genetics: Why does this take a centre stage in the genetics of microorganisms?
- Cell study: How this complex assessment has progressed.
- What tools can scientists wield in cell study?
- A look into the DNA of viruses.
- What can the COVID-19 virus help us to understand about genetics?
- Examining molecular genetics through chemical properties.
- Examining molecular genetics through physical properties.
- Is there a way you can store genetic information?
- Is there any distinction between molecular levels and subcellular levels?
- Variability and inheritance: What you need to note about living things at the molecular level.
- The research and study on molecular genetics: Key takeaways.
- What scientists can do within the confines of molecular genetics?
- Molecular genetics research and experiments: What you need to know.
- What is molecular genetics, and how can you learn about it?
Human Genetics Research Topics
Human genetics is an interesting field that has in-depth content. Some topics here will jog your brain and invoke curiosity in you. However, if you have difficulty writing a scientific thesis , you can always contact us for help.
- Can you extend your life by up to 100% just by gaining more understanding of the structure of DNA?
- What programming can you do with the help of DNA?
- Production of neurotransmitters and hormones through DNA.
- Is there something that you can change in the human body?
- What is already predetermined in the human body?
- Do genes capture and secure information on someone’s mentality?
- Vaccines and their effect on the DNA.
- What’s the likelihood that a majority of people on earth have similar DNA?
- Breaking of the myostatin gene: What impact does it have on the human body?
- Is obesity passed genetically?
- What are the odds of someone being overweight when the rest of his lineage is obese?
- A better understanding of the relationship between genetics and human metabolism.
- The truths and myths engulfing human metabolism and genetics.
- Genetic tests on sports performance: What you need to know.
- An insight on human genetics.
- Is there any way that you can prevent diseases that are transmitted genetically?
- What are some of the diseases that can be passed from one generation to the next through genetics?
- Genetic tests conducted on a person’s country of origin: Are they really accurate?
- Is it possible to confirm someone’s country of origin just by analyzing their genes?
Current Topics in Genetics
A list to help you choose from all the most relevant topics:
- DNA-altering experiments: How are scientists conducting them?
- How important is it to educate kids about genetics while they’re still in early learning institutions?
- A look into the genetics of men and women: What are the variations?
- Successes and failures in the study of genetics so far.
- What does the future of genetics compare to the current state?
- Are there any TV series or science fiction films that showcase the future of genetics?
- Some of the most famous myths today are about genetics.
- Is there a relationship between genetics and homosexuality?
- Does intelligence pass through generations?
- What impact does genetics hold on human intelligence?
- Do saliva and hair contain any genetic data?
- What impact does genetics have on criminality?
- Is it possible that most criminals inherit the trait through genetics?
- Drug addiction and alcohol use: How close can you relate it to genetics?
- DNA changes in animals, humans, and plants: What is the trigger?
- Can you extend life through medication?
- Are there any available remedies that extend a person’s life genetically?
- Who can study genetics?
- Is genetics only relevant to scientists?
- The current approach to genetics study: How has it changed since ancient times?
Controversial Genetics Topics
Last, but definitely not least, are some controversial topics in genetics. These are topics that have gone through debate and have faced criticism all around. Here are some you can write a research paper about:
- Gene therapy: Some of the ethical issues surrounding it.
- The genetic engineering of animals: What questions have people raised about it?
- The controversy around epigenetics.
- The human evolution process and how it relates to genetics.
- Gene editing and the numerous controversies around it.
- The question on same-sex relations and genetics.
- The use of personal genetic information in tackling forensic cases.
- Gene doping in sports: What you need to know.
- Gene patenting: Is it even possible?
- Should gene testing be compulsory?
- Genetic-based therapies and the cloud of controversy around them.
- The dangers and opportunities that lie in genetic engineering.
- GMOs and their impact on the health and welfare of humans.
- At what stage in the control of human genetics do we stop to be human?
- Food science and GMO.
- The fight against GMOs: Why is it such a hot topic?
- The pros and cons of genetic testing.
- The debates around eugenics and genetics.
- Labelling of foods with GMO: Should it be mandatory?
- What really are the concerns around the use of GMOs?
- The Supreme Court decision on the patent placed on gene discoveries.
- The ethical issues surrounding nurses and genomic healthcare.
- Cloning controversial issues.
- Religion and genetics.
- Behavior learning theories are pegged on genetics.
- Countries’ war on GMOs.
- Studies on genetic disorders.
Get Professional Help Online
Now that we have looked at the best rated topics in genetics, from interesting to controversial topics genetics, you have a clue on what to choose. These titles should serve as an example of what to select.
Nonetheless, if you need help with a thesis, we are available to offer professional and affordable thesis writing services . Our high quality college and university assignment assistance are available to all students online at a cheap rate. Get a sample to check on request and let us give you a hand when you need it most.

Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Comment * Error message
Name * Error message
Email * Error message
Save my name, email, and website in this browser for the next time I comment.
As Putin continues killing civilians, bombing kindergartens, and threatening WWIII, Ukraine fights for the world's peaceful future.
Ukraine Live Updates

- Vision, Mission and Purpose
- Focus & Scope
- Editorial Info
- Open Access Policy
- Editing Services
- Article Outreach
- Why with us
- Focused Topics
- Manuscript Guidelines
- Ethics & Disclosures
- What happens next to your Submission
- Submission Link
- Special Issues
- Latest Articles
- All Articles
- FOCUSED RESEARCH TOPICS

- Services Paper editing services Paper proofreading Business papers Philosophy papers Write my paper Term papers for sale Term paper help Academic term papers Buy research papers College writing services Paper writing help Student papers Original term papers Research paper help Nursing papers for sale Psychology papers Economics papers Medical papers Blog

126 High Quality Genetics Research Topics You Can Use

Most students look for genetics research topics to write about when pursuing biology studies. Developing exciting titles in this subject isn’t everyone’s favorite. Perhaps, that’s because of the technicality of this subject. This article lists interesting genetics topics for learners at different educational levels to consider for their papers. This list is essential because many learners struggle to develop or find titles for their essays but end with ideas that don’t interest educators.
What Is Genetics?
Genetics is the study of genes. In this field, scientists study how individuals pass down genes and traits from one generation to another. Genes carry vital information that impacts an individual’s appearance, health, and personality.
When writing research papers about genetics topics, students should follow several steps to create winning pieces. Here’s how to write an excellent essay in this field.
Pick a topic: Select a topic you’re comfortable researching, analyzing, and writing about to provide relevant and accurate information. Find sources and gather information: Look for reliable information sources, including peer-reviewed articles, journals, and professional books. Also, you can use websites with reliable information about genetics. Gather as much relevant information and analyze it. Outline the paper: Develop a structure to guide your research and writing. A typical outline should have an introduction, body, and conclusion. Write the paper: Use the outline and information from your research to write the essay. Ensure that your ideas and information flow logically.
The first and most vital step is selecting a good topic. Unfortunately, some learners have difficulties picking titles for their papers. Fortunately, the following list has a topic idea you will likely enjoy exploring. Use our online research paper writing servic e and get your paper done fast.
Interesting Topics In Genetics
Perhaps, you’re looking for an exciting title for your genetics paper. If so, this category has an ideal you will find exciting to explore.
- How does genetic variation impact the evolution of species?
- How do genes influence behavior and development?
- What role do genes play in disease susceptibility?
- How does the environment interact with genes to influence phenotype?
- How can we use genetics to improve crop plants or livestock?
- What is the impact of genetic engineering on society?
- How will the human genome project impact medicine?
- What ethical considerations are there in genetics research?
- How can we use genetics to predict individual risk for disease?
- What are the implications of personalized medicine?
- How accurate is genetic engineering?
- The bioethical and legal aspects of custom medicine based on the genetic composition
- The diagnostic challenge of newborns with heritable protein C deficiency
- Epithelial polarity in mammals and flies
- Risk factors contributions and medical care to trends in cardiovascular mortality
- Does cloning limit or increase biological diversity
- Understanding oculopharyngeal muscular dystrophy- is it an under-diagnosed disease?
- Imputation-based analysis- Quantitative traits and candidate regions
- Loss-of-function variants- A survey of human protein-coding genes
- Exploring DNA structural motifs’ thermodynamics
Explore any of these titles in your research, and your educator will find your paper exciting to read.
Controversial Topics In Genetics
Most people find controversial genetics topics exciting to read. Therefore, you can capture your educator’s attention by choosing and investigating a controversial idea in this study field. Here are such ideas to consider.
- Can animal cloning lead to health problems for humans?
- Is conducting maternal spindle transfer ethical?
- Is pronuclear transfer possible without causing mitochondrial disease in the embryo?
- Who decides the typical traits and which constitutes a disorder or disability?
- What are the ethical implications of improving human qualities like intelligence, height, and athletic ability?
- Is it ethical for doctors to alter germline traits using gene therapy?
- How honest are gene therapy’s protocol guidelines?
- Is changing a particular gene’s regulation ethical?
- Gene therapy ethics when curing genetic disease in a fetus
- How CRISPR-Cas9 gene editing affects a person’s germ line
- The ethics of genetic engineering
- The impact of genetic engineering on society
- The potential for abuse with genetic engineering
- The safety of genetically modified foods
- The regulation of genetic engineering
- The effect of genetic engineering on the environment
- The future of genetic engineering
- Is genetics the basis of depression?
- Can addictive substances change human genes?
- Can humans beat aging by altering genes?
These are controversial topics genetics students can find worth exploring. Nevertheless, prepare to research extensively to compose a winning paper about any of these ideas.
Human Genetics Topics For Research Papers
Human genetics entails studying inheritance in human beings. Here are some of the exciting human genetics topics for research papers.
- The Human Genome Project and its significance for understanding human genetics.
- Chromosomal abnormalities and their effects on human health.
- The role of genes in human development and behavior.
- The ethical implications of genetic testing and engineering.
- The impact of new technologies on the study of human genetics.
- The potential uses of genetic information in the diagnosis and treatment of disease.
- The social and economic implications of genetic discrimination.
- The role of genetics in predicting individual risk for common diseases.
- The impact of advances in genomic research on our understanding of human evolution.
- The role of genetics in human identity and individuality
- The link between human gene changes and diseases
- Genes coordination in human development
- How a fertilized egg directs the entire organism’s formation
- How gene editing to fix gene defects affects a human being
- How genetic disorders impact the heart’s pathological development
- Heritable genetic changes’ role in cardiovascular genetics
- How somatic mutations enhance tumor metastases
- How inherited genetic changes affect a person
- Human body processes that affect RNA and DNA sequencing
- How genetic mutations disrupt the normal cell proliferation regulation occurs in cancer
Consider these exciting human genetics topics if you find this sub-field exciting. Nevertheless, take the time to find sufficient and relevant information to write a comprehensive paper.
Molecular Genetics Topics
Molecular genetics is a biology sub-field addressing how different DNA molecule structures manifest in variations among organisms. Here are molecular genetics research paper topics to consider for your project.
- Disseminating superior genetics into a commercial population
- Analyzing linked genetic markets causing phenotypes differences
- Exploring various genes’ responses to environmental stressors
- How computer simulation affects molecular genetics
- Macromolecules that are vital in biological inheritance
- Molecular biology application in DNA forensics
- How the hereditary mechanism discovery impacted molecular genetics
- Tracing the molecular genetics origins from the 1930s
- Discussing double-helical structure in DNA molecule
- Ways of producing several copies of a DNA piece in the lab
- What more can humans possibly learn about DNA?
- How DNA determines the body structure
- DNA and terminal illness- Is there a connection?
- Does NDA sequencing have room for more?
- Describe and outline the latest molecular cancer genetics developments
- Explain genetic factors that enhance cancer susceptibility
- Are bacteria a genetic system?
- DNA and heredity- What’s the connection?
- Describe the shortcomings and potentials of stem cells
- Molecular techniques- Analyzing RNA, DNA, and proteins
Please select one of these titles and develop it into an exciting paper. Nevertheless, prepare to research extensively to fill your report with valuable information.
Current Topics In Genetics
Genetics research has a fascinating landscape with many current topics ripe for exploration. Here are some of the exciting contemporary ideas for research in genetics.
- Man versus bat’s molecular structure
- Genomics companies chasing after IPO- What are the impacts?
- 5G technology and how it affects the human genome
- Exploring the human microbiome evolution
- RNA binding and its role in leukemia treatment
- Analyzing the double-stranded RNA
- RNA-binding proteins characteristics
- Exploring the potential use of gene editing in treating COVID-19
- How drugs development and genes study relate
- How social engineering affects genetics
- Differentiating non-ethical and ethical gene therapy
- Gene therapy could make it for the rich people
- Why do people do gene testing under false names or anonymously?
- Insurers should force individuals to undergo testing
- Does euthanasia apply in the case of diseases?
- Who should know about different genetic disorders?
- Is genetic screening an interference with a person’s privacy?
- Ethical implications of newborn’s prenatal screening
- Can gene therapy treat a disease?
- Can gene therapy make people not accept people that are different?
These are good genetics paper topics for learners at various educational levels, especially if they want to write about the latest development in this field.
Hot Topics In Genetics
Perhaps, while thinking “I must do my term paper” you’re looking for topics that most people will want to read about and understand your viewpoint. In that case, here are genetics research paper topics to consider.
- Investigating critical molecules in genetic bone disease and bone development
- Intracellular traffic jams in genetics- Studying the multi-organellar
- Substance dependence and genetics- Is there a connection?
- Hereditary ovarian and breast cancers- Exploring preventive measures and causes
- Exploring gene composition in breast cancer and estrogen metabolism pathway
- Discuss cancer genetics and the cycle of the Eukaryotic cell
- Critical research on genetic elements that scientists can transport
- Genetic challenges in human diseases and RNA metabolism
- Genetic factors that cause human type 1 diabetes
- Approaching the relationship between human obesity and genetic susceptibility
- Is growing human organs morally upright and ethical?
- Human cloning science- Exploring its development and history
- Drug addiction and gene alteration
- Are genetically modified foods safe for animal and human consumption?
- Exploring fetus and genetic diagnosis
- DNA structure analysis from the genetic perspective
These are hot titles to explore in research. But like those in the other section, students must research each idea extensively to write high-quality papers. Contact us with a “ do my research paper ” request and get a winning paper done by professional writer.
Genetics Topics For Presentation
The genetics subject can be complex and broad. Therefore, choosing unique topics in genetics can benefit the learners’ presentations. Here are sample topics to consider for your presentation.
- The Genetic impact of neurological and terminal diseases
- Genetic engineering- What are the pros and cons
- Can humans create an artificial gene using synthetic chromosomes?
- Will cloning follow genetic engineering and research?
- The significance and complexity of gene mutation
- The advantages and unlimited potential of human genetics
- What does an individual’s DNA analysis say about their genetics?
- Is conducting genetic testing necessary?
- Is genes’ patent ownership ethical?
- Can humans isolate and eliminate hereditary conditions with genetic research?
This list has at least one example of a topic in this study field that you might want to explore. However, pick a title you will comfortably work with and deliver a quality paper that will prompt the educator to award you the top grade.
Get Professional Help With Research Paper
Maybe you have a title but no time to write a winning paper. Perhaps you’ve realized that you need thesis help to compose medical term papers that will impress the educator and earn you the desired grade. In that case, contact us for the best-rated research paper assistance. We offer affordable writing services to learners across educational levels. Contact us now for cheap but quality online help with your research project.

Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
Terms & Conditions Loyalty Program Privacy Policy Money-Back Policy
Copyright © 2013-2024 MyPaperDone.com
Genetics Research Paper Topics

- Biotechnology
- Birth defects
- Clone and cloning
- Genetic disorders
- Genetic engineering
- Human Genome Project
- Mendelian laws of inheritance
- Nucleic acid
The History of Genetics
Humans have known about hereditary characteristics for thousands of years. That knowledge has been used for the improvement of domestic plants and animals. Until the late nineteenth century, however, that knowledge had been gained by trial-and-error experiments. The modern science of genetics began with the pioneering work of the Austrian monk and botanist Gregor Mendel (1822–1884).
Academic Writing, Editing, Proofreading, And Problem Solving Services
Get 10% off with 24start discount code.
Mendel studied the genetic characteristics of pea plants. He was interested in finding out how certain traits, such as flower color and plant height, were passed on from generation to generation. During his lifetime, he studied dozens of generations of plants of all sizes, shapes, and colors. As a result of his research, Mendel was able to state a few basic laws describing the way genetic traits are inherited. He also came to the conclusion that there must be a specific biological unit responsible for the transmission of genetic traits. He called that unit a factor. Mendel’s “factors” were later given the name of genes.
Without question, Mendel was the father of the modern science of genetics. One of the great ironies of history, however, was that his discoveries were lost for more than three decades. Then, in the early 1900s, Mendel’s research was rediscovered almost simultaneously by three different biologists, the Dutch botanist Hugo de Vries (1848–1935), the German botanist Karl F. J. Correns (1864–1933), and the Austrian botanist Erich Tschermak von Seysenegg (1871–1962).
Although interest in genetics grew rapidly after 1900, a fundamental problem remained. Geneticists based all of their laws, theories, and experiments on the concept of the gene. But no one had any idea as to what was a gene. It seemed clear that the gene was probably some kind of chemical compound, or some combination of compounds. But no one had been able to determine exactly what kind of compound or compounds it was.
The answer to that question came in 1953. The American biologist James Watson (1928– ) and the English chemist Francis Crick (1916– ) collaborated to discover that a gene was a section of a very large and complex molecule found in the nuclei of all cells, the deoxyribonucleic acid (DNA) molecule.
The Chemistry of Genes
Imagine a very long chain of beads strung together to form a strand containing hundreds of thousands of beads. The strand contains beads of only four colors: red, yellow, blue, and green. That strand of beads can be compared to half of a DNA molecule. The other half of the molecule is a second strand almost identical to the first strand.
Watson and Crick showed that the sequence in which various colors of beads occur is significant. A DNA molecule in which the beads are arranged in the sequence blue-yellow-yellow-red-red-blue-blue-blue-, and so on, has meaning for a cell. The sequence tells the “chemical machinery” of the cell to make a certain kind of protein, such as the protein responsible for red hair or blue eyes. Another sequence of colors, for example, red-red-yellow-green-blue-green-red-, and so on, might be the “code” for making blonde hair or green eyes.
The components of a DNA molecule are not, of course, colored beads. They are certain groups of atoms known as nucleotides. Each nucleotide in a DNA molecule is comparable to one of the colored beads in the analogy above. Just as there are only four colors of beads in the above analogy, so there are only four different nucleotides in DNA molecules. Those nucleotides might be represented by the symbols A, C, G, and T (corresponding to bead colors of red, blue, green, and yellow). A DNA molecule, then, is a very long chain of nucleotides with a structure something like the following:
-C-T-A-T-C-G-A-C-T-T-G-A-C-T-T-T-G-C-C-A-C-A-A-C- …
The dots at the end of the chain indicate that the chain actually goes on much, much longer.
Watson and Crick said that each set of three nucleotides—they called them triads or codons—carried a specific message that cells could understand. Those messages told a cell to “make red hair,” or “make blue eyes,” or “help a person to grow tall,” or “give a person musical talent,” or any one of thousands of other traits that each human possesses.
This discovery answered the first question that geneticists had about heredity: how cells know which traits they are “supposed” to make and what functions they are “supposed” to carry out. The same discovery also answered the second question puzzling geneticists: how do these traits get passed down from generation to generation?
The answer to that question is that DNA molecules have the ability to make copies of themselves. When a cell divides (reproduces), so do the DNA molecules it contains. In most cases, two exactly identical molecules are produced from a single parent molecule.
When an egg cell (female reproductive cell) and a sperm cell (male reproductive cell) unite during fertilization, each cell provides DNA to the fertilized egg. The DNA from both parents combines to form DNA for the offspring. Whatever nucleotide sequences the mother and father had in their own cells, they pass on to their child.
Dominant and Recessive Traits
One fundamental question remains in the above example: suppose that a child is born to a father with red hair and a mother with blonde hair. What color hair will the child have?
Mendel worked with this question long before Watson and Crick discovered the nature of DNA. He found that for any one genetic trait, there were always two possible conditions. A flower might be red or white; a plant might be tall or short; a pea pod might be smooth or wrinkled; and so on. Mendel also discovered that one of these two conditions was more likely to “win out” over the other. He called the “winner” the dominant trait and the loser the recessive trait.
If a pea plant inherits a “tall” gene for height from both parent plants, the offspring is most like to be tall. If the pea plants inherits a “short” gene for height from both parent plants, the offspring is most likely to be short. But if the pea plant inherits a “tall” gene from one parent and a “short” gene from the second parent, the offspring is most likely to be tall.
An important part of Mendel’s work was finding out what the mathematical chances of various kinds of combinations might be. For example, he showed how to calculate the probabilities that would result when a “tall” parent pea plant was crossed with a “short” parent pea plant in the first, second, and succeeding generations.
The Future of Genetics
One can apply the principles of genetics in a great many situations without knowing anything about the structure of DNA molecules. However, the Watson-Crick discovery made possible a revolutionary change in the basic nature of genetics. As long as scientists had no idea as to what a gene was, there was not much they could do to make changes in the genes of a plant, animal, or human. But Watson and Crick showed that genes are nothing other than chemical compounds. If someone can make changes in chemical compounds in a laboratory, that person can also make changes in a DNA molecule. The problems faced are a good deal more difficult since DNA molecules are far more complex than most molecules that chemists work with. But the basic principles involved are the same.
Scientists are exploring a variety of ways in which genes can be modified to produce cells that can do things they could not do before. For example, it is possible to create the gene for the hormone (chemical messenger) known as insulin in a chemical laboratory. The work is fairly difficult, but by no means impossible. It simply requires that the correct atoms be assembled in the correct sequence. That artificial gene can then be inserted into the DNA of other organisms, such as bacteria. When the artificial gene becomes part of the bacterial DNA, it begins to function just like all the other genes in the bacteria’s DNA. The bacteria begins to function as an “insulin factory,” making a vitally important compound that it could never make before.
One of the most exciting recent developments in genetics is the initiation of the Human Genome Project, which officially began on October 1, 1990. This project is designed to provide a complete genetic road map outlining the location and function of the approximately 50,000 genes in human deoxyribonucleic acid (DNA) and to determine the sequences of the 3,000,000,000 base pairs that make up human DNA. As a result, genetic researchers will have easy access to specific genes to study how the human body works and to develop therapies for diseases. Gene maps for other species of animals also are being developed.
There appears to be virtually no technical limit to the things that scientists can do with genes. But with the promise of genetic research, many ethical and philosophical questions arise. One question is, of course, whether there are social or ethical limits to the kinds of changes scientists ought to be allowed to make in the genes of plants, animals, and humans. With research focusing on the ability to manipulate genes, there is the fear that the results will not always be beneficial. For the most part, the benefits for medicine and agriculture seem to far outweigh the possible abuses, and genetic research continues.
Back to Science Research Paper Topics .
ORDER HIGH QUALITY CUSTOM PAPER

341 Genetics Essay Topic Ideas & Examples
🏆 best genetics topic ideas & essay examples, 💡 most interesting genetics topics to write about, ✅ good research topics about genetics, ⭐ interesting topics to write about genetics, 🔎 genetics writing prompts, ✍️ genetics essay topics for college, 📌 simple & easy genetics essay titles, 👍 good essay topics on genetics, ❓ genetics essay questions.
- Genetically Modified Food Essay In spite of the perceived benefits of genetic engineering technology in the agricultural sector, the production and use of genetically modified foods has triggered a number of issues pertaining to safety and consequences of consumption.
- Genetically Modified Organisms: Views on GMOs For the reason that I was interested in GMOs and did my research before, the article did not change my perception of it much since I have already known what GMOs are and that they […] We will write a custom essay specifically for you by our professional experts 808 writers online Learn More
- The role of genetics in development In this case, the dominant gene will win over the recessive gene, and the child may exhibit the characteristics of a parent who produced dominant genes.
- Genetic Disorders: Causes and Treatment The individual inherits some of the characteristics from the mother and the rest is inherited from the father. Genetic disorders may be passed from the parents to the offspring’s during the process of fertilization.
- Danville Airlines’ Genetic Testing and Screening For instance, it may lead to the discrimination of the individual in the workplace. This implies that it was appropriate for Danville to conduct the tests with Reiger’s consent.
- Bisexuality: Genetics and Effects of the Sexual Behavior A homosexual is a person who is attracted to people of the same gender either physically, emotionally or sexually. Heterosexuals are people who are attracted physically, emotionally, and sexually to people of the opposite sex.
- Yeast Genetics and Complementation Lab Report In this study, the crossing of two mutants led to the development of phenotypic traits which are nothing but the colors, red or cream.
- Achondroplasia Genetic Disorder: Pedigree The pedigree problem is generally featured with the necessity to provide the correct connections among the family members in a genetic history chart.
- Should Parents Be Allowed to Choose the Characteristics of Their Children Through Genetic Manipulation? At the outset, genetic manipulation might be important to many parents as it trims down the prospects of grave infections in the newborn babies. The disadvantages of parents going for genetic manipulation seem to outweigh […]
- The Genetic Algorithm: Automatic Examination Timetable Scheduling Phenotype, on the other hand, represents the population in the search space corresponding to reality or a representation of the solutions through corresponding absolute values.
- Genetic and Genomic Technology Positive results mean that a patient has been diagnosed with the disease, and so treatment is essential to ensure the patient’s good health.
- Religious vs Scientific Views on Genetic Engineering With the need to increase the global economy, the field of agriculture is one among the many that have been used to improve the commercial production to take care of the global needs for food […]
- Should All Genetically Modified Foods Be Labeled? According to this scholar, members of the public are always comfortable with the idea of not labeling the genetically modified food.
- Agriculture and Genetics Disciplines Relationship The collapse of Crick’s theory was a setback to the genetics discipline because the foundations of genetic engineering are based on the central dogma premise.
- Genetics’ Role in Healthcare of Patents This paper focuses on genetics role in healthcare of patents and defines the language of genetic manipulation, its safety, legal and ethical issues, as well as mandatory screening and the role of the healthcare providers […]
- Genetic Manipulation of Human Embryos: Bioethical Issues Nonetheless, although the modification of human genotype may help in achieving a perfect genetic composition and eliminate a number of genetically transmitted diseases, there is a looming risk. The assembling of genetic makeup to enhance […]
- Privacy in Genetic Testing and Restriction of Access to Own Personal Information In a workplace context, genetic testing is associated with the most obvious intrusion of privacy within the aspects of job insurance coverage and promotion.
- Comparison of Theories of Addiction: The Biological Model and the Genetic Model Genetic and biological models aim at disclosing the essence of addiction as something natural and irreversible and the methods which are supported by neurobiology and physiology and become more appropriate for using and controlling human […]
- Mendelian Corn Genetics: An Experiment Seeds are then sorted out on the basis of their color and shape and the obtained data recorded adjacent to the respective phenotypes. Determine the 2 value for each experiment, and use the table of […]
- Marfan Syndrome in Genetic Counseling The two generation hierarchies above and one generation hierarchy below the Anne’s generation was pooled and presented in the chart as below: Firstly, the typical clinical symptoms attributed to MFS were sorted from the description […]
- The Study of Genetics: Importance for Society A key advantage of genetics for the clinical environment is the real possibility of manipulating the genetic code of a patient’s DNA in order to edit it.
- Concept of Genetic Cross Among Drosophila Melanogaster The hypothesis of the experiment was that the second generation of flies could be used to determine the phenotypes of both groups depending on the results of the experiment.
- Down Syndrome as the Most Common Genetic Condition in the US Firstly, to describe Down syndrome and the life of people with this disorder, it is necessary to give a scientific definition to this condition and underline the causes. People with Down syndrome are also people, […]
- Human Genetic Engineering: Key Principles and Issues There are many options for the development of events in the field of genetic engineering, and not all of them have been studied. To conclude, human genetic engineering is one of the major medical breakthroughs, […]
- Influence of Genetic Factors on Personality Heritability of personality is one of the most contentious issues in the field of modern psychology. Overall, the use of general personality characteristics in the analysis of twins compromises the reliability of evidence.
- Mitochondrial Diseases Treatment Through Genetic Engineering Any disorders and abnormalities in the development of mitochondrial genetic information can lead to the dysfunction of these organelles, which in turn affects the efficiency of intracellular ATP production during the process of cellular respiration.
- Preimplantation Genetic Diagnosis and Manipulation The promise of such disease reduction measures may inspire propositions from minorities in ECOSOC to advocate for PGD’s accessibility and investment in research.
- Aspects of the Genetic Enhancement Genetic enhancement means using genetic editing technologies to introduce changes into the genome of the fetus to achieve improvements in the physical or mental health of the future child.
- Patenting of Genetic Information Completing the sequencing of the nucleic acid sequence of the human genome led to the mass patenting of genes in the United States.
- Genetic Engineering: Is It Ethical to Manipulate Life? In the case of more complex operations, genetic engineering can edit existing genes to turn on or off the synthesis of a particular protein in the organism from which the gene was taken.
- Cystic Fibrosis: A Comprehensive Overview of the Genetic Causes and Pathophysiology In the development of cystic fibrosis, three main points are leading: lesions of the external secretion glands, changes in the connective tissue, and water and electrolyte disorders.
- Genetic and Environmental Impact of the Chornobyl Disaster The ecological impact of the explosion on the lands surrounding Chornobyl comes first. Chornobyl remains the worst in human history due to radioactive contamination.
- Genetics and Antipsychotics: Usefulness of Pharmacogenetic Analysis The response to antipsychotics can be observed as complex phenotypes, a combination of genetic and clinical elements, and symptoms that vary in severity as well as in adherence level.
- Crop Genetic Erosion: Understanding and Responding to Loss of Crop Diversity The recombining and shuffling of genes are expected to create more complexity and increase the capacity of the species to survive in the changing world.
- Can the Human Race Survive Without Genetically Modified Food? Although genetically modified food is a recent invention, the humankind will be unable to survive without it due to the rise in the global population.
- Children’s Temperament and Personality: Genetic Basis Biology is seen to play a major role in influencing children’s temperaments and personality traits, with outcomes being a result of genetic determination.
- The Morality of Prenatal Genetic Screening Most of the time, “genetic screening has been more associated with this option in the collective mental, rather than the possibility to better address a specific condition, leading to the complex discussion of an ethical […]
- Genetic Technology Integrated in Modern Society The recombinant DNA technique has a variety of uses and has enabled the development of novel enzymes that are well-suited for usage in particular food processing settings.
- Genetically Modified Organisms: Benefit or Harm? In other words, scientists may choose the DNA of the foods that some individuals may be allergic to, which can be harmful if they eat GMO crops.
- Experimentation in Inheritance and Genetics Eventually, a solution to the problem of development not referring to the cytoplasm was reached. In this regard, it is evident that experimentation plays a central part in the history of inheritance and genetics.
- Breast Cancer as a Genetic Red Flag It is important to note that the genetic red flags in Figure 1 depicted above include heart disease, hypertension, and breast cancer.
- T. Dobzhansky’s Input to Synthesis in Genetics One of the chapters titled “Dobzhansky, Waddington, and Schmalhausen: Embryology and the Modern Synthesis” discusses his views on the evolutionary theories of Schmalhausen and Waddington.
- Role of Nutritional Genetics in Health In the recent past, there has been a heightened interest in applying genomic technologies to comprehend the role of diet in disease development and health.
- Historical Development of Embryology and Epigenetics The theory of preformationism was widely recognized from the late 17th to the end of the 18th century. This concept proposed the occurrence of the generation of offspring due to the unfolding and development of […]
- Genetic Screening: Advantages and Disadvantages The results of screening can be used for monitoring a patient, the prevention of occurrence of certain diseases, and the treatment of diseases after diagnosis.
- Ethical Analysis: Preimplantation Genetic Diagnosis Furthermore, it is imperative to ascertain the goals of the therapy and the probability of achieving success. The last step in this model of decision-making is acting.
- Epigenetics: How Twin Mice Can Look Different In contrast to genetic changes, epigenetic shifts can be reversed because they have no impact on the DNA sequence; however, they can impact the way in which one’s organism reads the sequence.
- Genetic Mutation and Noonan Syndrome In general, the more nucleotide sequences that are impacted by a change, the more significant the impact of the conversion and the greater the likelihood that the mutation would be harmful.
- Genetically Modified Organisms (GMOs) in Food Production Thus, in case of GMOs, it is necessary to acknowledge the internal motivations and character in people that adhere to this type of food production.
- Aspects and Characteristics of Epigenetics In addition, the value of this source is that it shows the relationship between epigenetics and the occurrence of abnormalities such as diabetes and obesity.
- Genetic Counseling, Its Role, and Candidates In such cases, the benefits of such testing can be better explained to enable other family members to be tested and determine any other possible genetic problems.
- International Bioethics and Genetics Genetic discrimination is a problem of bioethical significance in which a patient’s confidential rights are violated to create favorable conditions on the part of the person or company who is the subject of the discriminatory […]
- The Ethical Dimensions of Genetic Testing If there is a hereditary condition that runs in the family, a DNA test can detect the presence of such conditions.
- Breast Cancer: Genetics and Malignancy In the presence of such conditions, the formation of atypical cells is possible in the mammary gland. In the described case, this aspect is the most significant since it includes various details of the patient’s […]
- Genetically Modified Organisms: Ethical Perspective Of course, some use the deontological approach and state that it is simply wrong to interfere with genetic codes as it is the divine domain.
- The Ethics of Genetic Modification In particular, the discussion of genetic modifications circulates around the possibilities of human embryo modifications and ethical considerations of the consequences of such experiments.
- Researching the Concept of Epigenetics When it comes to the considerations of epigenetics in terms of a disease for which an individual is at high risk, it is necessary to consider family history as well as environmental factors that add […]
- Genetics and Genomics in Healthcare Development The fourth dwells on the global utilization of genetics and genomics research while the fifth is an analysis of the influence of various factors on the utilization of genomics and genetics in healthcare.
- Genetic Disease in a Pregnant Woman and Fetus The patient should consider the relationship between oncogenesis and pregnancy and consider folic acid to maintain birth defects. Folic acid is a common supplement during pregnancy to make new cells and prevent the development of […]
- Triplet Nature of the Genetic Code One of the assumptions of molecular biology from an evolutionary perspective is that the triplet nature of the genetic code is a traditional form of two-nucleotide coding.
- Epigenetics: Analysis of Article Based on the completed family history assessment, I would not wholly link my risk metric to the outcome of the investigation and infer that I am vulnerable to the above-identified conditions. The outcome could be […]
- Idea of “Designer Babies” and Genetic Manipulations To date, the idea of “designer babies,” which claims that it is possible to alter the genes of the embryo, carrying out specific genetic manipulations, is becoming pretty popular but needs to be explored more.
- The Role of Genetics and Diet of Acne in Teenagers It is significant that the number of relapses, the duration of the course of therapy, and the increase in the number of patients with moderate and severe forms of acne directly depend on the adherence […]
- Examination of Albinism Genetic Disease In the majority of albinos, the lack of melanin leads to such symptoms as the absence of pigmentation in skin and hair.
- Gene Therapy and Genetic Enhancement On the other hand, genetic enhancement targets modifying the genes to augment the aptitudes of an organism outside the ordinary. Somatic gene editing impacts the cells of an individual under treatment and it is inherited […]
- The Reasons for Genetic Counseling According to Abacan, “genetic counseling is the process of helping people understand and adapt to the medical, psychological and familial implications of genetic contributions to disease”.
- Kidney Stones and Patient’s Genetics and Epigenetics Citrate inhibits the development of kidney stones through the formation of calcium citrate complexes thereby preventing the formation of insoluble crystals.
- Cloning: Genetically Identical Copy The clone develops in the womb and eventually, the adult female gives birth, with the new clone having an identical genetic makeup to the organism from which the somatic cell originated.
- Epigenetics: Subject and Example of Study Epigenetics refers to the study of how behaviors and the environment can cause changes that affect the way genes work. They are not static in the human body and are dynamically altered by changes in […]
- Down Syndrome Genetics and Behaviors Using current research literature on behavioral issues and novel treatments for Down syndrome, this paper explores and discusses behavioral inflexibility, restrictive and repetitive behaviors, and Down syndrome’s neurogenetic nature.
- Genetics: Only Girls Born in Miejsce Odrzanskie, Poland However, it may also be necessary to understand the limitations of such a study and consider the possibility of a coincidence.
- Genetic Testing: Advantages and Disadvantages At the same time, I acknowledge all the benefits that genetic testing can bring in terms of diagnosing a wide range of diseases and conditions.
- Mitosis and Genetic Makeup of Different Species As the centromeres of a cell align among the spindle equator, the genetic material of the maternal cell is duplicated, which allows for the two daughter cells to emerge.
- ZIP Code Prevails Over Genetic Code One of the health determinants is diet, which depends directly on the climate a person lives in and the scope of food products they can afford.
- Genetic Modification and Cloning Even though it is hard to predict all the outcomes of genetic modification and cloning, I would suggest using CRISPR Cas9 in treating retinal diseases such as the one described in the case study.
- Color Blindness and Its Genetic Nature Nevertheless, color blindness genes may be carried by the non-color-blind female and transferred to future generations. Depending on the mutation, inherited color blindness may be congenital or may reveal itself in childhood or adulthood.
- Genetic Modifications of Human and Animal Species As for the genetic modification of animals and insects, it can also be beneficial. In the case of humans, there must also be clear boundaries for modification.
- Environmental and Genetic Factors That Influence Health The factors of inaccessibility of proper treatment, poor lifestyle, and environmental agents’ exposure combined with the costs of addressing the CKD challenge lead to disparities in populations.
- SCN8A-Related Epilepsy – Genetic Seizure Disorder The paper contains the discussion of the standardized procedure for this diagnosis, suggests how the present experience would affect the medical practice concerning this kind of epilepsy.
- Molecular Genetics: Gene Sequence Homology The emergence of the Mendelian genetics in the 19th century and the discovery of DNA structure by James Watson and Francis Crick in the 20th century have paved the way for the development of molecular […]
- Genetic Testing: Screening for Colon Cancer This disorder is characterized by the development of hundreds of thousands of adenomatous polyps in the colon and rectum early in life.
- DNA Profiling and Required Genetic Testing The reliable tests for conducting genetic testing should be more than one in order to remove the element of doubt on matching DNA bands.
- The Relationship Between Epigenetics and the Effects of the Holocaust Tests are most likely to identify existing changes of DNA and the proteins related to DNA, which are responsible for the structure of the DNA and the availability of other elements related to the DNA.
- Acute Myeloid Leukemia: Genetic Features of Black Patients According to the researcher, the differences in the biological impact of disease and the socioeconomic factors play a crucial role in the disparity between the Blacks and the Whites in the recovery process.
- Resolving Genetic Issues or Playing With Mother Nature? Because mother nature is essential in sustaining human life, it is not right to play with mother nature. It is not right to play with mother nature because of the consequences of playing with her.
- Genetically Modified Food: Health Risks The main research question of the future study for me as a person with 1st Degree in Food and Nutrition will be the question of the harm of eating genetically modified foods and the possible […]
- Barlow’s Syndrome: Genetics of Mitral Valve Prolapse and Its Clinical Impact For the patients with mitral valve prolapse experiencing the episodes of tachycardia and rapid heartbeats the use of beta-blockers is allowed. The patient should be allowed to share their concerns and feelings about the mitral […]
- The Development of the Neural System and Genetic Program In the process of determining the connections worth keeping, a person’s brain takes into account their lived experiences and daily life, which in turn shape the direction of a person’s neural growth.
- Genetic Disorder: “A Genetic Link to Anorexia” The author effectively proves that the development of anorexia nervosa may occur not only due to the exposure to the social pressure of beauty standards, but also the presence of a genetic predisposition.
- Genetic Enhancement: Ethical Aspect In addition, it can take the shape of cosmetic modifications, which change the overall basis of human uniqueness and the unalterable aspect of the human body.
- Genetic Modification and Implicit Bias Against People With Disabilities There is also a factor of disabilities that are life-threatening to a child, or illnesses that may be able to be fatal within the first few years of life.
- Biotechnology and Genetic Engineering Apart from that, there are some experiments that cannot be ethically justified, at least in my opinion, for example, the cloning of human being or the attempts to find the gene for genius.
- Genetics and Genomics in Healthcare The first phase was used to determine the suitability of the survey tool while the main survey led to the collection of the desired data.
- Genetics, Reproductive and Cloning Technology in “Frankenstein” If Mary Shelley was for the idea of cloning technology, I think her novel would have ended up with Frankenstein creating a female companion for the monster to compliment the theme of love in the […]
- Genetic Mapping in the United States Genetic mapping is allowed and regulated by the Genetic Information Non-discrimination Act in the United States. On the contrary, genetic mapping of children in the United States can be conducted with the consent of a […]
- Knowing One’s DNA Genetic Makeup: Pros and Cons In addition, the knowledge that one might not get a job or insurance because of their genetic makeup is stressful and depressive.
- Genetics of Human Social Behavior Twin study method addresses questions such as what effects do environmental factors have on the twin similarities and twin differences.study which stands for Genome wide association is a method that seeks to establish the components […]
- The Factors That Cause Instances of Genetic Diversity Genetic diversity is a term that is used to refer to the difference in characteristics that occurs among members of the same species.
- Genetic Testing: Should You Bother to Exercise? However, some people are of the view that the effect of genetic variation on exercise-mediated body responses is dismal. This argument is usually based on the idea that several other factors such as diet, lifestyle, […]
- Genetic Screening for Mandatory Obesity Screening of Children The progress in behavioral genetics points to the significance of biological heredity in character traits, as well as in medical conditions. A good example is in the case of cancer where the affected genetic condition […]
- Genetically Modified Food: Analysis and Implications A section of scientists are opposed to the idea with claims that it brings about irreversible damage to biodiversity by changing the natural setting of the environment. Genetic engineering involves alteration of the cell which […]
- Epigenetics of High Fructose Corn Syrup at a Molecular Level Increasing or decreasing the amount of glucose concentration level in the blood, directly affects the concentration of fructose in blood, since they all act as determinants of the overall blood concentration In this case, high […]
- Genetic Engineering in the Movie “Gattaca” by Niccol This would not be right at all since a person should be responsible for their own life and not have it dictated to them as a result of a societal construct created on the basis […]
- Modern East Asians and Denisovans Share Genetic Material The researchers did not explain the specific mode of delivery of the genetic material through hybridization thus it must be assumed that the Denisovans in Siberia were able to travel to Southeast Asia and intermarry […]
- Genetic Counseling – Tay Sachs Disease In this case, there is a 25% likelihood of passing the gene to their children. This would be effective in preventing further passing down of the disease to their offspring.
- Psychiatric Genetics. Epigenetics and Disease Pathology The switching on and off of the imprinted genes is the same regardless of the parental origin. The genome-wide DNA analysis revealed that there was a difference in DNA methylation of the glucocorticoid receptor gene […]
- The Genetics of Crime: ‘Criminal Gene’ The idea that criminal and offending behavior stands in the correlation with the genetic features of the offender is not a novelty of our time.
- Genetic Family Historical Analysis In the family, Andrew is the only member who thinks that his disease is caused by a genetic predisposition. The above implies that Andrew should work closely with his physicians to ensure his therapy is […]
- Genetic Counseling Analysis To take a detailed family history, I would start with gathering the information about the consumers. Finally, I would ask about the members of the family who have already passed away and clarify the cause […]
- Genetics and the Asthma Case The allergies she complains of are some of the symptoms associated with asthma. Asthma is also known to attack children below the age of 15 years.
- Genetic Engineering Using a Pglo Plasmid The objective of this experiment is to understand the process and importance of the genetic transformation of bacteria in real time with the aid of extrachromosomal DNA, alternatively referred to as plasmids.
- A Genetic Clue to How Limbs Evolved From Fins The purpose of the study is to unveil the mechanisms of the evolution and change of fin to limb. The researchers used a morphological analysis of fossils and skeletons of modern animals, which supports the […]
- Genetic Inheritance and Its Role in Obesity This essay therefore analysis the different formations of obesity, the causes and in particular the significance of inheritance in the occurrence of obesity.
- Managing Diabetes Through Genetic Engineering Genetic engineering refers to the alteration of genetic make-up of an organism through the use of techniques to introduce a new DNA or eliminate a given hereditable material. What is the role of genetic engineering […]
- Genetic Diseases: Sickle Cell Anemia This genetic disorder research paper aims to elucidate the underlying molecular causes of SCA as well as its symptoms, inheritance, treatment, diagnosis, and prevalence in certain populations.
- Genetically Identical Twins and Different Disease Risk The study of MZ twins offers fascinating insights that help researchers explore the link between the sequence of the genotype and the phenotype.
- Genetic Disorders That Can Be Treated With Gene Therapy It is in this context that the application of gene therapy has increased the hope of medical professionals in overcoming and controlling such failures in the treatment of genetic disorders.
- Using Genetically-Modified Bacteria to Fight Cancer at Johns Hopkins To do so, a concise summary of the article will be provided, followed by a review of its relevance to the course.
- Diabetes Mellitus Type 2: The Family Genetic History This paper aims at analyzing family genetic history of a family, evaluating the impact of the family history on an adult participant’s health and planning a future wellness change to promote the wellness of the […]
- Forecast of Genetic Technology Therefore, this essay forecasts the advancement of genetic technology in the 22nd century in aspects of gene therapy and eugenics with the view of assessing its potential benefits and dangers.
- Genetic Male Pattern Baldness Vertex hair loss: Vertex hair loss can be observed on the top of the head which is crown area and does not touch the hairline of the forehead.
- Watching Television: Genetic Influence This can be testified by the manner the research question correlates to the dynamics of examining why watching TV has such a huge influence on genetics.
- Museum Genetic Presentation When this condition is violated, the population is opened allowing individuals to move from one population to another hence creating a net flow of genes which results to genetic variations and consequently, to evolution.
- Genetic Information in the Trosack Case The purpose of this essay is to integrate genetic information in the Trosack case to ensure that the patients can deal with the genetic basis of the disease in their child.
- The Genetic Basis of Human Cancer This is one of the most difficult in curing, as it may affect any part of the body, and seriously damage the body tissues.
- Advanced Pathophysiology: Genetic Technology In accordance to Tay-Sachs disorder, the specialist is likely to provide the following information: the origin of the disorder, what factors contribute to the occurrence of the disorder, characteristics of the disorder, treatment if available […]
- Effects of the Interaction Genetic Diversity This study though is trying to show the synergism between the UV-B and low genetic diversity as possible proof of the hypothesis that the UV-B could be the possible cause of declining amphibian populations.
- Biological and Genetic Influences on Criminality Men are twice as likely to be the victim of an assault or a robbery and 50 percent more likely to experience some crime of theft.
- Genetics of Prostate Cancer and Physical Features However, even though observations attest to the highly hereditary nature of the disease, research in the field so far has proved to be inconclusive, with most scientists failing to isolate the gene or genes which […]
- Criminal Justice and DNA: “Genetic Fingerprinting” DNA is one of the popular methods used by criminologists today, DNA technique is also known as “genetic fingerprinting”.the name given the procedure by Cellmark Diagnostics, a Maryland company that certified the technique used in […]
- Genetic Basis of Fitness Differences in Natural Populations In the article to summarize, the authors recognized that one way genomics affect biology is the possibility of identifying and studying how the characteristics affecting fitness, a key issue in natural selection, are genetically based.
- Computational Modelling and Genetic Regulatory Networks Analysis in Development The studies also include the control of transcription and the mechanisms of RNA maturation. The key determinants of cell fate are the transcription factors.
- Biotechnology and Animal Welfare: How Genetically Modified Chicken Serves the Demand in Fast Food Chains Beef was the most often used meat for the restaurants due to its containing in burgers, however, in 2020, the tendency started to move in the direction of chicken consumption.
- Breast Cancer Risk Factors: Genetic and Nutritional Influences However, the problems of genetics contribute to the identification of this disease, since the essence of the problem requires constant monitoring of the state of the mammary glands to detect cancer at an early stage.
- Breast Cancer Genetics & Chromosomal Analysis In this paper, the chromosomal analysis of breast cancer will be assessed, and the causes of the disorder will be detailed.
- The Role Genetics Information Plays in Treating Cancer Another process that also causes the disease to develop is called gene amplification and refers to the situation of the emergence of multiple copies of the same gene.
- Ethical Issues on Genetic Modified Baby: CRISPR-Ca9 Genetic Modification The discussion presented below gives a detailed analysis of the subject of genetically modified babies and the ethical values associated with the entire scientific process.
- Biotechnology, Genetics and Reproduction On the one hand, this is an opportunity to become parents for infertile couples, on the other hand, the ART industry acts as a new type of business and, therefore, we can talk about the […]
- Leukemia Types: Characteristics, Genetics, and Symptoms Leukemia is widely referred to as a group of blood cancers and is classified by the type of white blood cells and by the rate of disease progression over time.
- Evolutionary Biology: Program Model at Genetic Level The constant development of new technologies, natural sciences, and ideological factors causes a change in perceptions of the human body’s model and the impact of social processes on the behavior of the individual.
- Genetics of Sexual Orientation: Privacy, Discrimination, and Social Engineering The study tested the DNA of 400 gays and established a section of the X chromosome called Xq28. The outcome of the research is not limited to the research team only.
- Genetics as a Field and Its Practical Use Even in newborn screening, an area where genetic testing is excelling, parents opt to terminate the pregnancy for lack of a better solution to their condition.
- Clinical and Genetic Aspects of Neurofibromatosis Neurofibromatosis is an autosomal dominant genetic disorder that is caused by a mutation of a gene and characterized by a growth of tumors in nerves which may affect the structure of bones and skin conditions.
- Genome: Bioethics and Genetic Engineering Additionally, towards the end of the documentary, the narrator and some of the interviewed individuals explain the problem of anonymity that is also related to genetic manipulations.
- Researching the Genetic Enhancement: Unethical Practice and Social Norms One of the challenges that have emerged with the advent of genetic enhancement is the inability to ensure that all people have access to the technology.
- Genetic Test’s Availability to Employers and Insurers The introduction of genetics has gone a long way in sparking a heated debate as to why such companies are finding it necessary to introduce it to the list of information that the assured must […]
- Genetically Modified Foods: Substantial Equivalence Main principle of this concept is that genetically modified foods should be considered safe and reliable as conventional foods if the nutritional quality and compositions of GM foods are same as conventional foods.
- The Interrelationships and Implications of Genetic Discoveries In addition to this, the new sub-discipline comes with the ability to illuminate controversial topics within the field of medical sociology such as stem cell research and analysis of the interrelationships in human embryo and […]
- The Debate Pertaining to Genetic Modification This article looks at the setting up of a body, Royal Commission on Genetic Modification or the acronym RCGM on May 2000 to collect views from both the members of the public and scientific experts […]
- Genetic Predisposition to Alcohol: The Appreciation and Therapy for Alcoholism Through family studies it has been established that the likelihood of alcohol dependence and similar complications happening is more in the families of the individuals who have been affected as compared to in the people […]
- Epigenetics Influence on Adopted Embryos The exciting news is the role of epigenetics or influence of the adoptive mother’s body has on the DNA of the embryo as it grows using the mother’s nourishment, energy, and systems.
- Mental Illness Around the World: Socio-Cultural and Genetic Aspects Discussing the relationship between socio-political concept of people’s cultural identity and the degree of their susceptibility to mental illnesses represents a certain challenge, especially given the fact that the etiology of such illnesses has not […]
- Genetically Modified Organisms: For and Against The fact is that, genetic modification is the changing of the DNA code by the means of the genetic engineering, thus, the genes of the organisms are deviating from the normal genes of similar organisms, […]
- Ethical Problem of Genetic Testing and Chromosomal Abnormalities The testing can also be done to diagnose the genetic disorder, detecting future risks of contracting diseases as a result of genetic disorders, how these disorders respond to drugs, and detecting the extent of risks […]
- Unlocking Human Genome: Genetic Testing and Screening In the case of 64 year old Joe Miller, I recognize and understand that the patient has the right to die with dignity and that that includes dying on his own terms.
- Genetics of Parkinson Disease-Associated PARK2 Gene The following description highlights various aspects of the Human Genome Project that are thought to be connected in a series of events in influencing the cloning of the PARK2 gene.
- Genetic Research and Related Promises & Concerns More than 4000 diseases are thought to result from the defective functioning of a gene or a set of genes and if it were possible to identify the location or site of each gene on […]
- Genetic Difference in Explaining Athletic Performance Thus, St Louis argues that science on race and sports is not viable as it limits the explanation to physiological factors ignoring the importance of social and economic variables.
- Criminal Behavior: Role of Environment and Genetics In the Information age where a person has access to more knowledge about the folly of being involved in criminal activities and the negative impact of having a prison record, it is a mystery why […]
- The Various Aspects of Genetic Research Even a slight change in the pairing of the nucleotides will completely change the behavior of the human in question. Also, the genetic engineering helps the scientist to find out what is the function of […]
- Anderson and Genetic Research, Evolutionism & Creationism
- Genes and Environment: Genetic Factors and Issues Analysis
- Genetic Engineering Is Ethically Unacceptable
- Genetic Analysis: Term Definition and Molecular Genetics Experiments
- Issues of Sharing a Patient’s Genetic Information
- Ethical Issues Involving Genetic Test Accounts
- Creating a Genetically Modified Organism
- Genetics: State of Otter Conservation
- Genetic Basis for Alcoholism
- Genetically Modified Potato as Hepatitis B Vaccine
- Genetic Predisposition to Breast Cancer: Genetic Testing
- Genetic Testing Under Americans With Disabilities Act
- Genetic Counseling Preventing Pulmonary Arterial Hypertension
- Genetic Screening of Newborns and Its Benefits
- Evolution of Limbs: Fossil and Genetic Information
- Understanding Genetically Modified Foods by Howard et al.
- Next Generation Sequencing in Genetics
- Cystic Fibrosis: Genetics and Gender Factor
- 23andMe, a Genetics and Health Company
- Crispr/Cas9 Impact on Medical Genetics
- Growing GMO Seeds: Monsanto Corporation
- Genetic Disorders: The Turner and Down Syndromes
- Genetically Modified Foods: Pros or Cons
- Why Human Genetics Is Important
- Plasmids, Their Characteristics and Role in Genetics
- Type 2 Diabetes From Cultural and Genetic Aspects
- Genetic Testing in the Neonatal Realm: Technique of Assessing a Predisposition
- Genetic Testing and Related Health Issues
- Genetics: the Eugenics Movement
- Genetically Modified Foods and Pesticides for Health
- Genetics: “Bad Blood” Educational Series by BBC
- Genetically Engineered Food Against World Hunger
- Genetic Mapping and Its Social, Ethical, and Legal Implications
- Genetic Factors of Huntington’s Disease Progression
- “Intelligence, Race, and Genetics” by Sternberg et al.
- Genetic and Social Bond Theories in Criminology
- Epigenetics and Its Role in Cancer Detection and Prevention
- The Role of Epigenetics in Cancer: Contributors to the Formation of Cancer Tumors
- Genetics: the Erroneous Concept of Blending Inheritance
- Genetically Modified Foods: Pros and Cons
- Genetic Factors of Speech Coding at the Subcortical Level
- Reproductive and Genetic Technology in Infertility Treatment
- Genetically Modified Seeds in Environmental Context
- Genetically Modified Foods: Scientific Resources
- Genetically Modified Organisms in Canadian Agriculture
- What Concepts Create Misconception in Genetics?
- Genetic Testing Limitation: Ethical Perspective as a Framework
- Genetically Modified Salmon Labeling Issues: Biotechnology, Religious Beliefs, and Eating Preferences
- Molecular Genetics and Biological Inheritance
- Thalassemia as a Genetic Disorder and Its Management
- Huge Ethical Concerns: Human Genetic Creation and the Bible
- Prostate Cancer, Its Genetics and Prevention Methods
- Biodiversity, Its Evolutionary and Genetic Reasons
- Genetic Testing & Counseling and Their Value
- Abortion: Quality of Life and Genetic Abnormalities
- Ethics of Genetic Testing: Screening and Monitoring
- Advanced Diagnostic Procedures: Genetic Screening Pros and Cons
- Evolutionary Theory and Genetics
- Froma Harrop Views on Genetically Modified Food
- Genetics and Sociological Theories
- Genetically Modified Organisms in Farming
- Nutrition: Is Genetically Modified Food Bad or Good?
- Natural Sciences: Genetics Processes
- GMO Production: Reasons and Potential Effects
- Genetically Modified Foods: Should They Be Consumed?
- Do Our Genes Determine Learning Ability?
- Benefits and Concerns Regarding Genetically Modified Crops
- The Genetics of Alcohol Dependence
- Genetic Experimentation and Development
- Genetic Modification and Testing: Ethical Considerations
- Schizophrenia Genetic and Environmental Factors
- Ecological Effects of the Release of Genetically Engineered Organisms
- In Vitro Fertilization and Pre-implantation Genetic Diagnosis
- Epidemiology: Genetics-Related Programs
- Advanced Diagnostic Procedures: The Individual Impact of Genetic Diagnosis
- Possible Benefits of New Genetics
- The Effect of Genetically Modified Food on Society and Environment
- Proposition 37 and Genetically Engineered Foods
- Genetically Modified Food of Monsanto Company
- How Do Genetic and Environmental Factors Contribute To The Expression of Depression?
- Genetically Modified Corn in the United States of America
- Genetic and Cultural Differences Are Not Two Opposites
- The Human Genome Project and Its Revolutionary Insight into the Genetic Blue Print of the Human Body
- The Underpinnings of Genetic Diversity – Contribution of Darwin and Vavilov
- Genetically Modified Food and European Consumer Behavior
- Genetically Modified Foods Projects
- The Extent of Genetic Experimentation and Developments
- Genetically Modified Organisms and Controversial Discussions in Australia
- Overview on the Effects of Genetically Modified Food
- Is It Ethical to Abort Based On Genetic Disability?
- The Biotechnology Importance in Genetic Modification
- Can Genetically Modified Food Feed the World: Agricultural and Biotechnological Perspective
- Genetically Modified Foods Negative Aspects
- Analyzing the Prospects of Genetically Modified Foods
- Will Genetically Modified Foods Doom Us All?
- The Specifics of Society Genetic Constitution
- Single Nucleotide Polymorphisms Genetic Epidemiology
- The Evolutionary Genetics of Mycobacterium Tuberculosis
- Genetically Modified Foods and Environment
- The Debate Pertaining to Genetically Modified Food Products
- Is Genetically Modified Food Safe for Human Bodies and the Environment?
- Consumer Judgment on Genetically Modified Foods
- Genetic Alteration of Food Sources: A New Strategy to Improve Food Production
- Business Ethics-Labeling Genetically Modified Food
- Objection to the Production of Genetically Modified Foods
- The Importance of Facial Attractiveness on Genetic Diversity
- Ethical Implication of Human Genetics Research
- Is Genetically Engineered Food the Solution to the World’s Hunger Problems?
- Towards Understanding the Causes of Genetic Diversity
- The Correlation Between Genetics and Environmental Lifestyle
- The Roles of Genetics and Nurture on People with Dyslexia
- Tourette Syndrome: Symptoms, Causes, and Genetics
- The Relationship between Genetics and Religion
- The Theory Of Inheritance Within The Field Of Genetics
- An Analysis of the Role of Genetics and Environment in Causing Alcoholism
- Understanding the Basics of Genetics and Its Diseases
- Nature, Nurture and Egalitarian Policy: What Can We Learn from Molecular Genetics
- Significance of Discoveries in Genetics and DNA
- Winter Wheat Breeding, Genetics, And Cultivar Development
- The Issue of Genetics and Intelligence in the Article All in the Genes
- The Use of Genetics in Insurance and Impliations
- The Contribution of Family History, Age and Genetics to the Development of Alzheimer’s Disease
- Masters Program With The Department Of Molecular Genetics
- The Development Of Genetics Food Modifying Techniques: Analyzing The Effect Of Continuing Development
- Why Genetics Is Important And A Huge Part Of Our Lives
- Molecular Genetics: Catching the Criminal Using Electrophoresis
- The Importance of Genetics and Individuality in the Formation of a Personality
- How Does Genetics Affect The Achievement Of Food Security
- The Inheritance of Economic Status: Education, Class, and Genetics
- An Overview of the Principles Behind the Genetics and the Biochemical Prosprects
- The Importance Of Mendel’s Laws In Modern Genetics
- The Role of Genetics and Social Structure to the Rising Rate of Homicide in Miami
- The Effects Of Genetics And Environment On Biofilm Growth
- The Ethical Issues Surrounding the Field of Genetics Technology
- The Major Scientific Breakthroughs of Introns and Exons in Genetics
- What Part Do Genetics Play In Autoimmune Diseases
- The Genetics, Structure, Function, And Regulation Of Alpha Amylase
- Ongoing Intelligence Debate and Considerations of Environment, Culture, and Genetics
- The Genetics of Addiction Hereditary or Learned Behaviour
- The New Genetics of Mental Illness by Edmund S. Higgins
- The Importance of Family, Community and Culture Over Genetics and Individual Characteristics in Outliers, a Novel by Malcolm Gladwell
- The Cause of Our Overall Fear and Its Link to Genetics and Evolution Process in Our Fear of Immigrants, an Article by Jeremy Adam Smith
- The Life and Work of Gregor Mendel, the Father of Genetics
- The Establishment of a State Fisheries Genetics Program in Illinois
- Politically Correct Fanatics Their Denial Of Patterns And Genetics Among People
- The Significance of Selective Engineering in Genetics
- Genetics And The Possible Causation Of Autism Spectrum Disorders
- The Evolutionary Factors That Have Shaped The Genetics
- The Engineering Of Human Genetics In Dreams And Nightmares
- The Public and Private Sectors in the Process of Innovation: Theory and Evidence from the Mouse Genetics Revolution
- The Aspects of the Connection between the Environment and Genetics
- An Analysis of Scientific Knowledge About Founder Mutations in Genetics
- James Watson and his Contributions DNA and Genetics
- The Causes of Aging in Humans and the Impact of Genetics and Lifestyle on Age
- Genetics: Alcoholism and Normative Developmental Trajectory
- Is Homosexuality A Personal Choice Or Is It Genetics
- Will Genetics Destroy Sports?
- How Can Drug Metabolism and Transporter Genetics Inform Psychotropic Prescribing?
- Are Gender Roles Defined by Society or by Genetics?
- How Did the Drosophilia Melanogaster Impact Genetics?
- Does Genetics Affect Childhood Obesity?
- How Does Genetics Affect the Achievement of Food Security?
- Can Crop Models Identify Critical Gaps in Genetics, Environment, and Management Interactions?
- How Does Genetics Influence Human Behavior?
- Does Genetics Matter for Disease-Related Stigma?
- How Did Dolly Sheep Change Genetics Forever?
- Is Genetics Responsible for Allergies?
- How Does Genetics Affect Caffeine Tolerance?
- Can Genetics Cause Crime or Are We Presupposed?
- How Does Genetics Affect Child Development?
- Is Genetics Responsible for Mental Illnesses?
- How Do Genetics and Environment Affect a Child’s Behaviors?
- Can Genetics Reveal the Causes and Consequences of Educational Attainment?
- How Do Genetics and the Environment Influence One’s Self-Identity?
- What’s Genetics Engineering?
- How Do Neuroscience and Behavioral Genetics Improve Psychiatric Assessment?
- Why Will Tampering With Our Genetics Be Beneficial?
- How Does the Environment Change Genetics?
- Will Benchtop Sequencers Resolve the Sequencing Trade-off in Plant Genetics?
- Why Is Genetics Important in Real Life?
- How Does Genetics Play a Role in Aging?
- Chicago (A-D)
- Chicago (N-B)
IvyPanda. (2024, February 26). 341 Genetics Essay Topic Ideas & Examples. https://ivypanda.com/essays/topic/genetics-essay-topics/
"341 Genetics Essay Topic Ideas & Examples." IvyPanda , 26 Feb. 2024, ivypanda.com/essays/topic/genetics-essay-topics/.
IvyPanda . (2024) '341 Genetics Essay Topic Ideas & Examples'. 26 February.
IvyPanda . 2024. "341 Genetics Essay Topic Ideas & Examples." February 26, 2024. https://ivypanda.com/essays/topic/genetics-essay-topics/.
1. IvyPanda . "341 Genetics Essay Topic Ideas & Examples." February 26, 2024. https://ivypanda.com/essays/topic/genetics-essay-topics/.
Bibliography
IvyPanda . "341 Genetics Essay Topic Ideas & Examples." February 26, 2024. https://ivypanda.com/essays/topic/genetics-essay-topics/.
- Genetic Engineering Topics
- DNA Essay Ideas
- Cloning Questions
- Bioethics Titles
- Experiment Questions
- Epigenetics Essay Titles
- Down Syndrome Topics
- Nature vs Nurture Research Topics
- Biomedicine Essay Topics
- Disorders Ideas
- Animal Ethics Research Ideas
- Discovery Ideas
- Disability Essay Topics
- Gender Inequality Research Topics
Research Topics & Ideas
Biotechnology and Genetic Engineering

If you’re just starting out exploring biotechnology-related topics for your dissertation, thesis or research project, you’ve come to the right place. In this post, we’ll help kickstart your research topic ideation process by providing a hearty list of research topics and ideas , including examples from recent studies.
PS – This is just the start…
We know it’s exciting to run through a list of research topics, but please keep in mind that this list is just a starting point . To develop a suitable research topic, you’ll need to identify a clear and convincing research gap , and a viable plan to fill that gap.
If this sounds foreign to you, check out our free research topic webinar that explores how to find and refine a high-quality research topic, from scratch. Alternatively, if you’d like hands-on help, consider our 1-on-1 coaching service .

Biotechnology Research Topic Ideas
Below you’ll find a list of biotech and genetic engineering-related research topics ideas. These are intentionally broad and generic , so keep in mind that you will need to refine them a little. Nevertheless, they should inspire some ideas for your project.
- Developing CRISPR-Cas9 gene editing techniques for treating inherited blood disorders.
- The use of biotechnology in developing drought-resistant crop varieties.
- The role of genetic engineering in enhancing biofuel production efficiency.
- Investigating the potential of stem cell therapy in regenerative medicine for spinal cord injuries.
- Developing gene therapy approaches for the treatment of rare genetic diseases.
- The application of biotechnology in creating biodegradable plastics from plant materials.
- The use of gene editing to enhance nutritional content in staple crops.
- Investigating the potential of microbiome engineering in treating gastrointestinal diseases.
- The role of genetic engineering in vaccine development, with a focus on mRNA vaccines.
- Biotechnological approaches to combat antibiotic-resistant bacteria.
- Developing genetically engineered organisms for bioremediation of polluted environments.
- The use of gene editing to create hypoallergenic food products.
- Investigating the role of epigenetics in cancer development and therapy.
- The application of biotechnology in developing rapid diagnostic tools for infectious diseases.
- Genetic engineering for the production of synthetic spider silk for industrial use.
- Biotechnological strategies for improving animal health and productivity in agriculture.
- The use of gene editing in creating organ donor animals compatible with human transplantation.
- Developing algae-based bioreactors for carbon capture and biofuel production.
- The role of biotechnology in enhancing the shelf life and quality of fresh produce.
- Investigating the ethics and social implications of human gene editing technologies.
- The use of CRISPR technology in creating models for neurodegenerative diseases.
- Biotechnological approaches for the production of high-value pharmaceutical compounds.
- The application of genetic engineering in developing pest-resistant crops.
- Investigating the potential of gene therapy in treating autoimmune diseases.
- Developing biotechnological methods for producing environmentally friendly dyes.
Biotech & GE Research Topic Ideas (Continued)
- The use of genetic engineering in enhancing the efficiency of photosynthesis in plants.
- Biotechnological innovations in creating sustainable aquaculture practices.
- The role of biotechnology in developing non-invasive prenatal genetic testing methods.
- Genetic engineering for the development of novel enzymes for industrial applications.
- Investigating the potential of xenotransplantation in addressing organ donor shortages.
- The use of biotechnology in creating personalised cancer vaccines.
- Developing gene editing tools for combating invasive species in ecosystems.
- Biotechnological strategies for improving the nutritional quality of plant-based proteins.
- The application of genetic engineering in enhancing the production of renewable energy sources.
- Investigating the role of biotechnology in creating advanced wound care materials.
- The use of CRISPR for targeted gene activation in regenerative medicine.
- Biotechnological approaches to enhancing the sensory qualities of plant-based meat alternatives.
- Genetic engineering for improving the efficiency of water use in agriculture.
- The role of biotechnology in developing treatments for rare metabolic disorders.
- Investigating the use of gene therapy in age-related macular degeneration.
- The application of genetic engineering in developing allergen-free nuts.
- Biotechnological innovations in the production of sustainable and eco-friendly textiles.
- The use of gene editing in studying and treating sleep disorders.
- Developing biotechnological solutions for the management of plastic waste.
- The role of genetic engineering in enhancing the production of essential vitamins in crops.
- Biotechnological approaches to the treatment of chronic pain conditions.
- The use of gene therapy in treating muscular dystrophy.
- Investigating the potential of biotechnology in reversing environmental degradation.
- The application of genetic engineering in improving the shelf life of vaccines.
- Biotechnological strategies for enhancing the efficiency of mineral extraction in mining.
Recent Biotech & GE-Related Studies
While the ideas we’ve presented above are a decent starting point for finding a research topic in biotech, they are fairly generic and non-specific. So, it helps to look at actual studies in the biotech space to see how this all comes together in practice.
Below, we’ve included a selection of recent studies to help refine your thinking. These are actual studies, so they can provide some useful insight as to what a research topic looks like in practice.
- Genetic modifications associated with sustainability aspects for sustainable developments (Sharma et al., 2022)
- Review On: Impact of Genetic Engineering in Biotic Stresses Resistance Crop Breeding (Abebe & Tafa, 2022)
- Biorisk assessment of genetic engineering — lessons learned from teaching interdisciplinary courses on responsible conduct in the life sciences (Himmel et al., 2022)
- Genetic Engineering Technologies for Improving Crop Yield and Quality (Ye et al., 2022)
- Legal Aspects of Genetically Modified Food Product Safety for Health in Indonesia (Khamdi, 2022)
- Innovative Teaching Practice and Exploration of Genetic Engineering Experiment (Jebur, 2022)
- Efficient Bacterial Genome Engineering throughout the Central Dogma Using the Dual-Selection Marker tetAOPT (Bayer et al., 2022)
- Gene engineering: its positive and negative effects (Makrushina & Klitsenko, 2022)
- Advances of genetic engineering in streptococci and enterococci (Kurushima & Tomita, 2022)
- Genetic Engineering of Immune Evasive Stem Cell-Derived Islets (Sackett et al., 2022)
- Establishment of High-Efficiency Screening System for Gene Deletion in Fusarium venenatum TB01 (Tong et al., 2022)
- Prospects of chloroplast metabolic engineering for developing nutrient-dense food crops (Tanwar et al., 2022)
- Genetic research: legal and ethical aspects (Rustambekov et al., 2023). Non-transgenic Gene Modulation via Spray Delivery of Nucleic Acid/Peptide Complexes into Plant Nuclei and Chloroplasts (Thagun et al., 2022)
- The role of genetic breeding in food security: A review (Sam et al., 2022). Biotechnology: use of available carbon sources on the planet to generate alternatives energy (Junior et al., 2022)
- Biotechnology and biodiversity for the sustainable development of our society (Jaime, 2023) Role Of Biotechnology in Agriculture (Shringarpure, 2022)
- Plants That Can be Used as Plant-Based Edible Vaccines; Current Situation and Recent Developments (İsmail, 2022)
As you can see, these research topics are a lot more focused than the generic topic ideas we presented earlier. So, in order for you to develop a high-quality research topic, you’ll need to get specific and laser-focused on a specific context with specific variables of interest. In the video below, we explore some other important things you’ll need to consider when crafting your research topic.
Get 1-On-1 Help
If you’re still unsure about how to find a quality research topic, check out our Research Topic Kickstarter service, which is the perfect starting point for developing a unique, well-justified research topic.

You Might Also Like:

Submit a Comment Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- Print Friendly
204 Genetics Research Topics & Essay Questions for College and High School
Genetics studies how genes and traits pass from generation to generation. It has practical applications in many areas, such as genetic engineering, gene therapy, gene editing, and genetic testing. If you’re looking for exciting genetics topics for presentation, you’re at the right place! Here are genetics research paper topics and ideas for different assignments.
🧬 TOP 7 Genetics Topics for Presentation 2024
🏆 best genetics essay topics, ❓ genetics research questions, 👍 good genetics research topics & essay examples, 🌟 cool genetics topics for presentation, 🌶️ hot genetics topics to write about, 🔎 current genetic research topics, 🎓 most interesting genetics topics.
- Advantages and Disadvantages of Genetic Testing
- Should Parents Have the Right to Choose Their Children Based on Genetics?
- The Potential Benefits of Genetic Engineering
- The Importance of Heredity and Genetics
- Genetically Modified Pineapples and Their Benefits
- Cause and Effect of Genetically Modified Food
- Simulating the Natural Selection and Genetic Drift
- Genetic and Social Behavioral Learning Theories Learning and behavioral habits in human beings can be influenced by social, environmental and genetic factors. Genetic theory describes how genes help in shaping human behaviors.
- Genetically Modified Food as a Current Issue GM foods are those kinds of food items that have had their DNA changed by usual breeding; this process is also referred to as Genetic Engineering.
- Genetic and Environmental Impacts on Teaching Work If students do not adopt learning materials and the fundamentals of the curriculum well, this is a reason for reviewing the current educational regimen.
- Link Between Obesity and Genetics Obesity affects the lives through limitations implemented on the physical activity, associated disorders, and even emotional pressure.
- Benefits of Genetic Engineering The potential increase of people’s physical characteristics and lifespan may be regarded as another advantage of genetic engineering.
- Ban on Genetically Modified Foods Genetically modified (GM) foods are those that are produced with the help of genetic engineering. Such foods are created from organisms with changed DNA.
- Human Genetics: Multifactorial Traits This essay states that multifactorial traits in human beings are essential for distinguishing individual characteristics in a population.
- Genetic and Environmental Factors Causing Alcoholism and Effects of Alcohol Abuse The term alcoholism may be used to refer to a wide range of issues associated with alcohol. Simply put, it is a situation whereby an individual cannot stay without alcohol.
- Genetic Testing and Privacy & Discrimination Issues Genetic testing is fraught with the violation of privacy and may result in discrimination in employment, poor access to healthcare services, and social censure.
- A Career in Genetics: Required Skills and Knowledge A few decades ago, genetics was mostly a science-related sphere of employment. People with a degree in genetics can have solid career prospects in medicine and even agriculture.
- Genetically Modified Organisms: Pros and Cons Genetically modified organisms are organisms that are created after combining DNA from a different species into an organism to come up with a transgenic organism.
- Genetic Engineering: Dangers and Opportunities Genetic engineering can be defined as: “An artificial modification of the genetic code of an organism. It changes radically the physical nature of the being in question.
- Behavioral Genetics in “Harry Potter” Books The reverberations of the Theory of Behavioral Genetics permeate the Harry Potter book series, enabling to achieve the comprehension of characters and their behaviors.
- Relation Between Genetics and Intelligence Intelligence is a mental ability to learn from experience, tackle issues and use knowledge to adapt to new situations and the factor g may access intelligence of a person.
- Does Genetic Predisposition Affect Learning in Other Disciplines? This paper aims to examine each person’s ability to study a discipline for which there is no genetic ability and to understand how effective it is.
- Genetic Engineering: Cloning With Pet-28A Embedding genes into plasmid vectors is an integral part of molecular cloning as part of genetic engineering. An example is the cloning of the pectate lyase gene.
- Ethical Concerns on Genetic Engineering The paper discusses Clustered Regularly Interspaced Short Palindromic Repeats technology. It is a biological system for modifying DNA.
- Family Pedigree, Human Traits, and Genetic Testing Genetic testing allows couples to define any severe genes in eight-cell embryos and might avoid implanting the highest risk-rated ones.
- Technology of Synthesis of Genetically Modified Insulin The work summarizes the technology for obtaining genetically modified insulin by manipulating the E. coli genome.
- GMO Use in Brazil and Other Countries The introduction of biotechnology into food production was a milestone. Brazil is one of the countries that are increasingly using GMOs for food production.
- Genetics of Developmental Disabilities The aim of the essay is to explore the genetic causes of DDs, especially dyslexia, and the effectiveness of DNA modification in the treatment of these disorders.
- Genomics, Genetics, and Nursing Involvement The terms genomics and genetics refer to the study of genetic material. In many cases, the words are erroneously used interchangeably.
- GMO: Some Peculiarities and Associated Concerns Genetically modified organisms are created through the insertion of genes of other species into their genetic codes.
- Genetics Seminar: The Importance of Dna Roles DNA has to be stable. In general, its stability becomes possible due to a large number of hydrogen bonds which make DNA strands more stable.
- Genetically Modified Foods and Their Impact on Human Health Genetically modified food has become the subject of discussion. There are numerous benefits and risks tied to consumption of genetically modified foods.
- Nutrition: Obesity Pandemic and Genetic Code The environment in which we access the food we consume has changed. Unhealthy foods are cheaper, and there is no motivation to eat healthily.
- Mendelian Genetics and Chlorophyll in Plants This paper investigates Mendelian genetics. This lab report will examine the importance of chlorophyll in plants using fast plants’ leaves and stems.
- DNA and the Birth of Molecular Genetics Molecular genetics is critical in studying traits that are passed through generations. The paper analyzes the role of DNA to provide an ample understanding of molecular genetics.
- The Concept of Epigenetics Epigenetics is a study of heritable phenotypic changes or gene expression in cells that are caused by mechanisms other than DNA sequence.
- Literature Review: Acceptability of Genetic Engineering The risks and benefits of genetic engineering must be objectively evaluated so that modern community could have a better understanding of this problem
- Impacts of Genetic Engineering of Agricultural Crops In present days the importance of genetic engineering grew due to the innovations in biotechnologies and Sciences.
- How Much Do Genetics Affect Us?
- What Can Livestock Breeders Learn From Conservation Genetics and Vice Versa?
- How Do Genetics Affect Caffeine Tolerance?
- How Dolly Sheep Changed Genetics Forever?
- What Is the Nature and Function of Genetics?
- What Are the Five Branches of Genetics?
- How Does Genetics Affect the Achievement of Food Security?
- Are Owls and Larks Different in Genetics When It Comes to Aggression?
- How Do Neuroscience and Behavioral Genetics Improve Psychiatric Assessment?
- How Does Genetics Influence Human Behavior?
- What Are Three Common Genetics Disorders?
- Can Genetics Cause Crime or Are We Presupposed?
- What Are Examples of Genetics Influences?
- How Do Genetics Influence Psychology?
- What Traits Are Influenced by Genetics?
- Why Tampering With Our Genetics Will Be Beneficial?
- How Genetics and Environment Affect a Child’s Behaviors?
- Which Country Is Best for Genetics Studies?
- How Does the Environment Change Genetics?
- Can Crop Models Identify Critical Gaps in Genetics, Environment, and Management Interactions?
- How Can Drug Metabolism and Transporter Genetics Inform Psychotropic Prescribing?
- Can You Change Your Genetics?
- How Old Are European Genetics?
- Will Benchtop Sequencers Resolve the Sequencing Trade-off in Plant Genetics?
- What Can You Study in Genetics?
- What Are Some Genetic Issues?
- Does Genetics Matter for Disease-Related Stigma?
- How Did the Drosophila Melanogaster Impact Genetics?
- What Is a Genetics Specialist?
- Will Genetics Destroy Sports?
- Autism Spectrum Disorder in Twins: Genetics Study Autism spectrum disorder is a behavioral condition caused by genetic and environmental factors. Twin studies have been used to explain the hereditary nature of this condition.
- Genetic and Genomic Healthcare: Nurses Ethical Issues Genomic medicine is one of the most significant ways of tailoring healthcare at a personal level. This paper will explore nursing ethics concerning genetic information.
- Is ADHD Genetically Passed Down to Family Members? Genetic correlations between such qualities as hyperactivity and inattention allowed us to define ADHD as a spectrum disorder rather than a unitary one.
- Alzheimer’s Disease: Genetic Risk and Ethical Considerations Alzheimer’s disease is a neurodegenerative disease that causes brain shrinkage and the death of brain cells. It is the most prevalent form of dementia.
- Environmental Impact of Genetically Modified Crop In 1996, the commercial use of genetically modified (GM) crop production techniques had increasingly been accepted by many farmers.
- Gene Transfer and Genetic Engineering Mechanisms This paper discusses gene transfer mechanisms and the different genetic engineering mechanisms. Gene transfer, a natural process, can cause variation in biological features.
- Genetics in Diagnosis of Diseases Medical genetics aims to study the role of genetic factors in the etiology and pathogenesis of various human diseases.
- The Morality of Selective Abortion and Genetic Screening The paper states that the morality of selective abortion and genetic screening is relative. This technology should be made available and legal.
- Environmental Ethics in Genetically Modified Organisms The paper discusses genetically modified organisms. Environmental ethics is centered on the ethical dilemmas arising from human interaction with the nonhuman domain.
- Detection of Genetically Modified Products Today, people are becoming more concerned about the need to protect themselves from the effects of harmful factors and to buy quality food.
- Genetically Modified Organisms Solution to Global Hunger It is time for the nations to work together and solve the great challenge of feeding the population by producing sufficient food and using fewer inputs.
- Restricting the Volume of Sale of Fast Foods and Genetically Modified Foods The effects of fast foods and genetically modified foods on the health of Arizona citizens are catastrophic. The control of such outlets and businesses is crucial.
- Researching of Genetic Engineering DNA technology entails the sequencing, evaluation and cut-and-paste of DNA. The following paper analyzes the historical developments, techniques, applications, and controversies.
- Genetically Modified Crops: Impact on Human Health The aim of this paper is to provide some information about genetically modified crops as well as highlight the negative impacts of genetically modified soybeans on human health.
- Genetic Engineering Biomedical Ethics Perspectives Diverse perspectives ensure vivisection, bio, and genetic engineering activities, trying to deduce their significance in evolution, medicine, and society.
- Down Syndrome: The Genetic Disorder Down syndrome is the result of a glandular or chemical disbalance in the mother at the time of gestation and of nothing else whatsoever.
- Genetic Modifications: Advantages and Disadvantages Genetic modifications of fruits and vegetables played an important role in the improvement process of crops and their disease resistance, yields, eating quality and shelf life.
- Genetics of Personality Disorders The genetics of different psychological disorders can vary immensely; for example, the genetic architecture of schizophrenia is quite perplexing and complex.
- Labeling of Genetically Modified Products Regardless of the reasoning behind the labeling issue, it is ethical and good to label the food as obtained from genetically modified ingredients for the sake of the consumers.
- Convergent Evolution, Genetics and Related Structures This paper discusses the concept of convergent evolution and related structures. Convergent evolution describes the emergence of analogous or similar traits in different species.
- Genetic Technologies in the Healthcare One area where genetic technology using DNA works for the benefit of society is medicine, as it will improve the treatment and management of genetic diseases.
- Are Genetically Modified Organisms Really That Bad? Almost any food can be genetically modified: meat, fruits, vegetables, etc. Many people argue that consuming products, which have GMOs may cause severe health issues.
- Type 1 Diabetes in Children: Genetic and Environmental Factors The prevalence rate of type 1 diabetes in children raises the question of the role of genetic and environmental factors in the increasing cases of this illness.
- Discussion of Genetic Testing Aspects The primary aim of the adoption process is to ensure that the children move into a safe and loving environment.
- The Normal Aging Process and Its Genetic Basis Various factors can cause some genetic disorders linked to premature aging. The purpose of this paper is to talk about the genetic basis of the normal aging process.
- Defending People’s Rights Through GMO Labels Having achieved mandatory labeling of GMOs, the state and other official structures signal manufacturers of goods about the need to respect customers’ rights.
- Medicine Is Not a Genetic Supermarket Together with the development of society, medicine also develops, but some people are not ready to accept everything that science creates.
- Epigenetics: Definition and Family History Epigenetics refers to the learning of fluctuations in creatures induced by gene expression alteration instead of modification of the ‘genetic code itself.
- Genetically Modified Organisms in Aquaculture Genetically Modified Organisms are increasingly being used in aquaculture. They possess a unique genetic combination that makes them uniquely suited to their environment.
- Genetic Modification of Organisms to Meet Human Needs Genetic modification of plants and animals for food has increased crop yields as the modified plants and animals have more desirable features such as better production.
- Discussion of Epigenetics Meanings and Aspects The paper discusses epigenetics – the study of how gene expression takes place without changing the sequence of DNA.
- Genetic Testing and Bill of Rights and Responsibilities Comparing the Patient Bill of Rights or Patient Rights and Responsibilities of UNMC and the Nebraska Methodist, I find that the latter is much broader.
- Genetically Modified Products: Positive and Negative Sides This paper considers GMOs a positive trend in human development due to their innovativeness and helpfulness in many areas of life, even though GMOs are fatal for many insects.
- Overview of African Americans’ Genetic Diseases African Americans are more likely to suffer from certain diseases than white Americans, according to numerous studies.
- Plant Genetic Engineering: Genetic Modification Genetic engineering is the manipulation of the genes of an organism by completely altering the structure of the organism.
- Genetically Modified Fish: The Threats and Benefits This article’s purpose is to evaluate possible harm and advantages of genetically modified fish. For example, the GM fish can increase farms’ yield.
- Genetic Linkage Disorders: An Overview A receptor gene in the human chromosome 9 is the causative agent of most blood vessel disorders. Moreover, blood vessel disorders are the major cause of heart ailments.
- Natural Selection and Genetic Variation The difference in the genetic content of organisms is indicative that certain group of organisms will stay alive, and effectively reproduce than other organisms residing in the same environment.
- The role of genes in our food preferences.
- The molecular mechanisms of aging and longevity.
- Genomic privacy: ways to protect genetic information.
- The effects of genes on athletic performance.
- CRISPR-Cas9 gene editing: current applications and future perspectives.
- Genetic underpinnings of human intelligence.
- The genetic foundations of human behavior.
- The role of DNA analysis in criminal justice.
- The influence of genetic diversity on a species’ fate.
- Genetic ancestry testing: the process and importance.
- Genetically Modified Foods: How Safe are they? This paper seeks to address the question of whether genetically modified plants meant for food production confer a threat to human health and the environment.
- The Genetic Material Sequencing This experiment is aimed at understanding the real mechanism involved in genetic material sequencing through nucleic acid hybridization.
- Genetically Modified Organisms in Human Food This article focuses on Genetically Modified Organisms as they are used to produce human food in the contemporary world.
- Genetics and Public Health: Disease Control and Prevention Public health genomics may be defined as the field of study where gene sequences can be used to benefit society.
- Genetic Disorder Cystic Fibrosis Cystic fibrosis is a genetic disorder. The clinical presentation of the disease is evident in various organs of the body as discussed in this paper.
- The Study of the Epigenetic Variation in Monozygotic Twins The growth and development of an organism result in the activation and deactivation of different parts due to chemical reactions at strategic periods and locations
- Human Genome and Application of Genetic Variations Human genome refers to the information contained in human genes. The Human Genome Project (HGP) focused on understanding genomic information stored in the human DNA.
- Genetic Alterations and Cancer The paper will discuss cancer symptoms, causes, diagnosis, treatment, side-effects of treatment, and also its link with a genetic alteration.
- Saudi Classic Aniridia Genetic and Genomic Analysis This research was conducted in Saudi Arabia to determine the genetic and genomic alterations that underlie classic anirida.
- What Makes Humans Mortal Genetically? The causes of aging have been studied and debated about by various experts for centuries, there multiple views and ideas about the reasons of aging and.
- Decision Tree Analysis and Genetic Algorithm Methods Application in Healthcare The paper investigates the application of such methods of data mining as decision tree analysis and genetic algorithm in the healthcare setting.
- Genetic Screening and Testing The provided descriptive report explains how genetic screening and testing assists clinicians in determining cognitive disabilities in babies.
- Neurobiology: Epigenetics in Cocaine Addiction Studies have shown that the addiction process is the interplay of many factors that result in structural modifications of neuronal pathways.
- Genetic (Single Nucleotide Polymorphisms) Analysis of Genome The advancement of the SNP technology in genomic analysis has made it possible to achieve cheap, effective, and fast methods for analyzing personal genomes.
- Darwin’s Theory of Evolution: Impact of Genetics New research proved that genetics are the driving force of evolution which causes the revision of some of Darwin’s discoveries.
- Genetic Tests: Pros and Cons Genetic testing is still undergoing transformations and further improvements, so it may be safer to avoid such procedures under certain circumstances.
- Case on Preserving Genetic Mutations in IVF In the case, a couple of a man and women want to be referred to an infertility specialist to have a procedure of in vitro fertilization (IVF).
- Race: Genetic or Social Construction One of the most challenging questions the community faces today is the following: whether races were created by nature or society or not.
- Huntington’s Chorea Disease: Genetics, Symptoms, and Treatment Huntington’s chorea disease is a neurodegenerative heritable disease of the central nervous system that is eventually leading to uncontrollable body movements and dementia.
- Genetics: A Frameshift Mutation in Human mc4r This article reviews the article “A Frameshift Mutation in Human mc4r Is Associated With Dominant Form of Obesity” published by C. Vaisse, K. Clement, B. Guy-Grand & P. Froguel.
- DNA Profiling: Genetic Variation in DNA Sequences The paper aims to determine the importance of genetic variation in sequences in DNA profiling using specific techniques.
- Genetic Diseases: Hemophilia This article focuses on a genetic disorder such as hemophilia: causes, symptoms, history, diagnosis, and treatment.
- Genetics: Gaucher Disease Type 1 The Gaucher disease type 1 category is a genetically related complication in which there is an automatic recession in the way lysosomes store some important gene enzymes.
- Genetic Science Learning Center This paper shall seek to present an analysis of sorts of the website Learn Genetics by the University of Utah.
- What Is Silencer Rna in Genetics RNA silencing is an evolutionary conserved intracellular surveillance system based on recognition. RNA silencing is induced by double-stranded RNA sensed by the enzyme Dicer.
- Cystic Fibrosis: Genetic Disorder Cystic fibrosis, also referred to as CF, is a genetic disorder that can affect the respiratory and digestive systems.
- Genetics or New Pharmaceutical Article Within the Last Year Copy number variations (CNVs) have more impacts on DNA sequence within the human genome than single nucleotide polymorphisms (SNPs).
- Genetic Disorders: Diagnosis, Screening, and Treatment Chorionic villus is a test of sampling done especially at the early stages of pregnancy and is used to identify some problems which might occur to the fetus.
- Research of Genetic Disorders Types This essay describes different genetic disorders such as hemophilia, turner syndrome and sickle cell disease (SCD).
- Genetic Mechanism of Colorectal Cancer Colorectal Cancer (CRC) occurrence is connected to environmental factors, hereditary factors, and individual ones.
- Isolated by Genetics but Longing to Belong The objective of this paper is to argue for people with genetic illnesses to be recognized and appreciated as personages in all institutions.
- Genetic Association and the Prognosis of Phenotypic Characters The article understudy is devoted to the topic of genetic association and the prognosis of phenotypic characters. The study focuses on such a topic as human iris pigmentation.
- PiggyBac Transposon System in Genetics Ideal delivery systems for gene therapy should be safe and efficient. PB has a high transposition efficiency, stability, and mutagenic potential in most mammalian cell lines.
- Advantages of Using Genetically Modified Foods Genetic modifications of traditional crops have allowed the expansion of agricultural land in areas with adverse conditions.
- Genetic Factors as the Cause of Anorexia Nervosa Genetic predisposition currently seems the most plausible explanation among all the proposed etiologies of anorexia.
- Bioethical Issues in Genetic Analysis and Manipulations We are currently far from a point where we can claim that we should be providing interventions to some and not others due to their genetic makeup.
- Personality Is Inherited Principles of Genetics The present articles discusses the principles of genetics, and how is human temperament and personality formed.
- The Effects of Genetic Modification of Agricultural Products Discussion of the threat to the health of the global population of genetically modified food in the works of Such authors as Jane Brody and David Ehrenfeld.
- Genetic foundations of rare diseases.
- Genetic risk factors for neurodegenerative disorders.
- Inherited cancer genes and their impact on tumor development.
- Genetic variability in drug metabolism and its consequences.
- The role of genetic and environmental factors in disease development.
- Genomic cancer medicine: therapies based on tumor DNA sequencing.
- Non-invasive prenatal testing: benefits and challenges.
- Genetic basis of addiction.
- The origins of domestication genes in animals.
- How can genetics affect a person’s injury susceptibility?
- Genetic Engineering in Food and Freshwater Issues The technology of bioengineered foods, genetically modified, genetically engineered, or transgenic crops, will be an essential element in meeting the challenging population needs.
- Genetic Engineering and Religion: Designer Babies The current Pope has opposed any scientific procedure, including genetic engineering, in vitro fertilization, and diagnostic tests to see if babies have disabilities.
- Op-ED Genetic Engineering: The Viewpoint The debate about genetic engineering was started more than twenty years ago and since that time it has not been resolved
- All About the Role of Genetic Engineering and Biopiracy The argument whether genetically engineered seeds have monopolized the market in place of the contemporary seeds has been going on for some time now.
- Genetic Engineering and Cloning Controversy Genetic engineering and cloning are the most controversial issues in modern science. The benefits of cloning are the possibility to treat incurable diseases and increase longevity.
- Biotechnology: Methodology in Basic Genetics The material illustrates the possibilities of ecological genetics, the development of eco-genetical models, based on the usage of species linked by food chain as consumers and producers.
- Genetics Impact on Health Care in the Aging Population This paper briefly assesses the impact that genetics and genomics can have on health care costs and services for geriatric patients.
- Concerns Regarding Genetically Modified Food It is evident that genetically modified food and crops are potentially harmful. Both humans and the environment are affected by consequences as a result of their introduction.
- Family Genetic History and Planning for Future Wellness The patient has a family genetic history of cardiac arrhythmia, allergy, and obesity. These diseases might lead to heart attacks, destroy the cartilage and tissue around the joint.
- Personal Genetics and Risks of Diseases Concerning genetics, biographical information includes data such as ethnicity. Some diseases are more frequent in specific populations as compared to others.
- Genetic Predisposition to Alcohol Dependence and Alcohol-Related Diseases The subject of genetics in alcohol dependence deserves additional research in order to provide accurate results.
- Genetically-Modified Fruits, Pesticides, or Biocontrol? The main criticism of GMO foods is the lack of complete control and understanding behind GMO processes in relation to human consumption and long-term effects on human DNA.
- Genetic Variants Influencing Effectiveness of Exercise Training Programmes “Genetic Variants Influencing Effectiveness of Exercise Training Programmes” studies the influence of most common genetic markers that indicate a predisposition towards obesity.
- Eugenics, Human Genetics and Their Societal Impact Ever since the discovery of DNA and the ability to manipulate it, genetics research has remained one of the most controversial scientific topics of the 21st century.
- Genetic Interference in Caenorhabditis Elegans The researchers found out that the double-stranded RNA’s impact was not only the cells, it was also on the offspring of the infected animals.
- Genetics and Autism Development Autism is associated with a person’s genetic makeup. This paper gives a detailed analysis of this condition and the role of genetics in its development.
- Genetically Modified Food Safety and Benefits Today’s world faces a problem of the shortage of food supplies to feed its growing population. The adoption of GM foods can solve the problem of food shortage in several ways.
- Start Up Company: Genetically Modified Foods in China The aim of establishing the start up company is to develop the scientific idea of increasing food production using scientific methods.
- Community Health Status: Development, Gender, Genetics Stage of development, gender and genetics appear to be the chief factors that influence the health status of the community.
- Homosexuality as a Genetic Characteristic The debate about whether homosexuality is an inherent or social parameter can be deemed as one of the most thoroughly discussed issues in the contemporary society.
- Why Is the Concept of Epigenetics So Fascinating? Epigenetics has come forward to play a significant role in the modern vision of the origin of illnesses and methods of their treatment, which results in proving to be fascinating.
- Epigenetics and Its Effect on Physical and Mental Health This paper reviews a research article and two videos on epigenetics to developing an understanding of the phenomenon and how it affects individuals’ physical and mental health.
- Genetic Counseling for Cystic Fibrosis Some of the inherited genes may predispose individuals to specific health conditions like cystic fibrosis, among other inheritable diseases.
- Medical and Psychological Genetic Counseling Genetic counseling is defined as the process of helping people understand and adapt to the medical, psychological, and familial implications of genetic contributions to disease.
- Patent on Genetic Discoveries and Supreme Court Decision Supreme Court did not recognize the eligibility of patenting Myriad Genetics discoveries due to the natural existence of the phenomenon.
- Genetic Testing, Its Background and Policy Issues This paper will explore the societal impacts of genetic research and its perceptions in mass media, providing argumentation for support and opposition to the topic.
- Genetically Modified Organisms and Future Farming There are many debates about benefits and limitations of GMOs, but so far, scientists fail to prove that the advantages of these organisms are more numerous than the disadvantages.
- Mitosis, Meiosis, and Genetic Variation According to Mendel’s law of independent assortment, alleles for different characteristics are passed independently from each other.
- Genetic Counseling and Hypertension Risks This paper dwells upon the peculiarities of genetic counseling provided to people who are at risk of developing hypertension.
- The Perspectives of Genetic Engineering in Various Fields Genetic engineering can be discussed as having such potential benefits for the mankind as improvement of agricultural processes, environmental protection, resolution of the food problem.
- Labeling Food With Genetically Modified Organisms The wide public has been concerned about the issue of whether food products with genetically modified organisms should be labeled since the beginning of arguments on implications.
- Diabetes Genetic Risks in Diagnostics The introduction of the generic risks score in the diagnosis of diabetes has a high potential for use in the correct classification based on a particular type of diabetes.
- Residence and Genetic Predisposition to Diseases The study on the genetic predisposition of people to certain diseases based on their residence places emphasizes the influence of heredity.
- Eugenics, Human Genetics and Public Policy Debates Ethical issues associated with human genetics and eugenics have been recently brought to public attention, resulting in the creation of peculiar public policy.
- Value of the Epigenetics Epigenetics is a quickly developing field of science that has proven to be practical in medicine. It focuses on changes in gene activity that are not a result of DNA sequence mutations.
- Genetically Modified Organisms: Position Against Genetically modified organisms are organisms that are created after combining DNAs of different species to come up with a transgenic organism.
- Genetically Modified Organisms and Their Benefits Scientists believe GMOs can feed everyone in the world. This can be achieved if governments embrace the use of this new technology to create genetically modified foods.
- Food Science and Technology of Genetic Modification Genetically modified foods have elicited different reactions all over the world with some countries banning its use while others like the United States allowing its consumption.
- How Much can We Control Our Genetics, at What Point do We Cease to be Human? The branch of biology that deals with variation, heredity, and their transmission in both animals and the plant is called genetics.
- Genetic Engineering: Gene Therapy The purpose of the present study is to discover just what benefits gene therapy might have to offer present and future generations.
Cite this post
- Chicago (N-B)
- Chicago (A-D)
StudyCorgi. (2022, January 16). 204 Genetics Research Topics & Essay Questions for College and High School. https://studycorgi.com/ideas/genetics-essay-topics/
"204 Genetics Research Topics & Essay Questions for College and High School." StudyCorgi , 16 Jan. 2022, studycorgi.com/ideas/genetics-essay-topics/.
StudyCorgi . (2022) '204 Genetics Research Topics & Essay Questions for College and High School'. 16 January.
1. StudyCorgi . "204 Genetics Research Topics & Essay Questions for College and High School." January 16, 2022. https://studycorgi.com/ideas/genetics-essay-topics/.
Bibliography
StudyCorgi . "204 Genetics Research Topics & Essay Questions for College and High School." January 16, 2022. https://studycorgi.com/ideas/genetics-essay-topics/.
StudyCorgi . 2022. "204 Genetics Research Topics & Essay Questions for College and High School." January 16, 2022. https://studycorgi.com/ideas/genetics-essay-topics/.
These essay examples and topics on Genetics were carefully selected by the StudyCorgi editorial team. They meet our highest standards in terms of grammar, punctuation, style, and fact accuracy. Please ensure you properly reference the materials if you’re using them to write your assignment.
This essay topic collection was updated on January 21, 2024 .
Thesis Helpers
Find the best tips and advice to improve your writing. Or, have a top expert write your paper.
150 Fantastic Genetics Research Topics To Write Best Thesis

Studies on aging, cancer, genetics, clinical research, or disease have a thing to do with genetics. Every medical student needs this, and it doesn’t matter if you just got admission to college, are in a postgraduate study, or you’re graduating from a medical school. Genetics paper topics or topics for your essays guide you in deciding which area to focus on for your paper.
You can’t write a paper or essay out of context, which is why a custom and reliable genetics topic list can help you understand what to write about to impress your professors and get the best grades. This blog post is all about that, with recommendations on how to write an excellent paper.
What Is Genetics?
On the other hand, genes are the basic unit of inheritance. They are a transportation medium that passes features and characteristics down from ancestors to offspring. For instance, eye and hair color is a solid example of gene inheritance. If your parents are redheads, there’s a higher chance that you’ll be too.
How To Write A Good Research Paper
A good research paper heavily depends on how effectively you do your research. Research helps you understand your topic, making it easier to convey your information to the necessary audience. These are some of the essential steps to writing your paper:
Understand your topic to be able to research and express your thoughts well. Do your research and write an outline Draw up your first draft after thorough research. Remember, first drafts aren’t always perfect. Revisit your introduction, body of text, and conclusion Revise and edit Create a checklist, and see if you cross-checked all aspects Refine your paper
These steps are necessary to create an excellent paper. You also need a creative topic to whip out a fantastic paper. These are 140 genetics paper topics you can consider.
Topics In Genetics
There are hundreds of genetics topics to explore and build on to develop a well-written paper. Here are 25 of them:
- Discuss the effects of genetics on human life.
- How does conservation genetics affect livestock breeding?
- Discuss the three general genetic disorders
- Can gene mutation aid clinical malaria treatment?
- Can genetics tests combat Alzheimer’s disease?
- Discuss the relationship between genes and living cells.
- Can human genetic formation be predetermined before birth?
- Combating HIV with genetic mutation; discuss the possibility and consequences.
- Can genetics help in treating intellectually disabled children?
- Discuss the examples of genetic influences on human.
- How does the environment influence human genes?
- Analyze the characteristics and traits that are solely caused by genetic inheritance
- Discuss the ethical implications of tampering with genetic imprints
- Is genetic build a prerequisite for sports players?
- How do genes affect mental health?
- Discuss the influence of genetics on obesity
- Is genetic mutation the anticipated cure for cancer?
- Examine the influence of genes on human behavior
- Thoroughly examine and discuss the importance of the five branches of genetics to the human body
- Discuss the relationship between food security and genetics in the world economy.
- Do genes determine a child’s character development?
- How do scientists decipher the genetic code?
- Examine the process of gene crossing in animals
- Discuss gene regulation in animals and plants
- Could genetics be the cure for numerous diseases?
- Discuss the influence of cloning on cancer treatment.
- Critically examine the structural syndrome and genetic influence of the long QT syndrome.
- Discuss the possibility of using genetic mutation to strengthen bone density and avoid bone fragility.
- Discuss the aftereffects of prenatal testing using diagnostic procedures
- Discuss the impact of sexual dimorphisms in biomarkers; address both the metabolic and the genetic biomarkers.
Interesting Genetics Topics
There are controversial genetics topics that are interesting in nature. If you want to write engaging research papers, here are 20 interesting topics you can consider:
- Discuss the relationship between DNA and Alzheimer
- The benefits of genetic tests; do they save lives?
- Analyze the impact of genetics on human aging
- Discuss the effect of genetics on human-like longevity.
- Analyze the unique characteristics that differentiate each human from the others.
- Discuss the percentage of parental genes transmission to offspring; which parent passes down more genes?
- Analyze the possibility of mental illness inheritance in relation to genetics
- Discuss the relationship between diets and genetics
- Examine the possibility of DNA modification
- What is the basis of mutation?
- Discuss the history of DNA change during human evolution
- Is it possible to change human genes?
- Discuss the nature of genetic influence on female
- Examine the possibility of successful cloning in the future
- Analyze the relationship between human genetics and allergies.
- Discuss the ethical perspectives of genetics regarding research processes.
- What are the current human genetics research methods?
- Discuss the possible discovery of Dinosaur DNA in modern science.
- Why do humans look different?
- Does genetics influence human behavior?
- Healthy living and genetics: discuss the better way to ensure long life-expectancy rate in humans
- Discuss the animals that match human DNA closely, and examine why the resemblance exists.
- Obesity in humans; can genetics be blamed?
- Is the male gender more immune to obesity?
- Discuss the preventive methods against general mental illnesses.
Genetics Topics For Research Papers
You need excellent genetics research paper topics to write great essays. You can find topics on molecular genetics, transmission, and population genetics for your papers here:
- Discuss the futuristic possibility of genetic cloning.
- Discuss the research extent and success of the biological dark matter of the human genome.
- Does gene inheritance influence human behavior in any way?
- Discuss the relationship between pediatric mental research and genetics
- Do gene characteristics affect your investing capabilities?
- Discuss the gorilla genome; how it helped rare apes escape extinction
- Does gene inheritance include asthma?
- Can genetic testing help patients with bone diseases?
- Can In-Vitro disrupt the genetic reproduction chain?
- The influence of genomic hybridization on fruits
- Examine the future of genetic coding
- Discuss the methods behind genetic engineering
- Discuss how genetics can improve an individual’s personality
- RNA — Give an analysis of the expression and its use during the creation of the COVID-19 vaccine
- What’s the role of clathrin function in developing new tricks for an old protein
- Complex challenges in genetics and public health
- Lender’s congenital amaurosis: what are the long-term consequences of gene therapy?
- Which framework is used to create a relative estimate of pathogenicity in human genetic variants?
- Osteoporosis: assess how genetic mutations help in the severe bone disease
- Human placentae: how is mosaicism distributed?
- Genetics engineering: what are the foremost principles to adhere to?
- How should doctors navigate using gene therapy in altering germline traits?
- Hyperechogenic kidneys: Explain the expected outcome of prenatally diagnosed kidneys
- Cancer vaccine: assess their efficacy based on quantitative data
- DNA modules: What’s the role of 3D printing?
Genetics Topics For Presentation
If you’re writing a class presentation, and are skeptical about the best topic to choose. These are ten superb genetics topics up for grabs:
- Discuss the science behind gene replacement
- Examine the impact of genetics on diseases
- Can genes form an immunity barrier against certain germs in humans?
- Examine the development of transgenic organisms in genetic research laboratories
- Can genetics be used to fight against Parkinson’s disease?
- Analyze and compare the animal DNA to humans, using high compatibility animals like cows in the process
- Is cloning the next genetic trend?
- Discuss the influence of epigenetics on cocaine addiction
- Are genetics the driving force of evolution?
- Discuss Cystic fibrosis as a genetic disorder
Controversial Topics In Genetics
Genetics holds several disputations among scientists. If you want a debatable paper that’ll introduce and solve controversies. Here are 15 controversial topics genetics branches offer you:
- Discuss the ethical beliefs surrounding gene therapy.
- Will cloning produce a negative outcome in the future?
- Examine the debate regarding artificial insemination and natural pregnancy
- Are genetic tests 100% accurate?
- Would you consider parent changing their children’s DNA before their birth an ethical practice?
- Should human organs be grown?
- Should genetic testing be conducted, or deemed unethical due to processes and outcomes?
- Discuss the controversy surrounding a bioethics revolution
- Discuss the religious beliefs on creating a perfect child
- Are patients’ data at stake during human genetics research?
- Explain the risk of discrimination that might come with genetics science
- Examine the hospital’s right to patent human genetics, with or without patients’ discretion.
- Discuss the advantages and disadvantages of human genome research
- Discuss the ethical implications of gene editing
- Can cloning cause lethal defects in humans and animals?
Hot Topics In Genetics
Numerous interesting topics in genetics are bound to tickle your fancy. If you want to write about the current medical situation in genetics, you can pick from any of these 15 topics:
- Are genetically modified foods safe for consumption?
- Discuss the genetic structure and symptoms of Huntington’s Chorea disease
- What are the implications of DNA research?
- Discuss the process of DNA alterations in plants
- What factors determine mutation in animals’ and plants’ DNA
- Can mental intelligence be genetically implanted?
- Discuss the unproven superstition about genetics
- Does genetics have any influence on homosexuality?
- What differentiates the genetic build of men and women?
- Why do women live longer than men?
- Is foundational genetics knowledge necessary in the high school curriculum?
- Will the human genome project do humanity good or bad?
- Discuss the process of RNA formation
- Can offspring inherit mutated genes?
- Discuss the science between blood groups.
Human Genetics Topics
Human genetics studies gene inheritance in humans. You can find research titles in areas like cytogenetics, genomics, clinical genetics, and genetics counseling here, as well as appropriate biology topics to write about . These topics will give you solid research ideas and a perfectly formulated paper:
- Discuss the kinds of genetically transmitted diseases
- Discuss the effects of myostatin deficiency in humans
- Discuss the predetermined characteristics of humans compared to those that can be changed.
- How to program with DNA
- Discuss the social and legal issues regarding genetic testing in your country
- Analyze the pharmacogenetics of alcohol abuse
- Provide insights into diseases that are discovered from family-based genomic studies.
- Discuss the implications of transmission genetics
- Explain the science behind the carpal tunnel syndrome
- Does genetic mutation affect reproduction in humans?
- Can you determine a person’s state of origin through genes?
- Discuss the prevention of genetically transmitted diseases
- Can DNA restructuring extend life expectancy?
- Discuss the controversy between human metabolism and genetics
- Do incestual relationships create malfunctioning genes?
- Discuss several genetic issues that plague the human body.
- Critically examine and explain the effect of Drosophila Melanogaster in genetics.
- Does genetic inheritance affect crime rates in a country?
- Discuss the negative impact of genetics on the human race
- Why are humans mortal?
Molecular Genetics Topics
Molecular genetics studies the structure of the DNA, its expression, and its replication, including its influence on the human body. You can choose any of these current topics in genetics to write a fantastic paper on molecular genetics;
- Discuss the experimental process of molecular genetics
- Examine the process involved in gene information storage
- Discuss the genetic formation and mutation in bacteria based on practical samples from any accessible laboratories
- Identify the differentiation factors at molecular and structural levels.
- Examine the simple and complex learning process of molecular genetics and how it contributes to the broader spectrum of the subject
- Examine the process of gene mutation
- Examine the study of the viral DNA
- Analyze the process of predicting potential prostate cancer with a polygenic score
- Discuss the possibility of paternal transmission of congenital myotonic dystrophy to an offspring.
- Discuss the gene mutation processes in breast cancer
Get Help With Genetics Research
The magic to writing well-constructed essays is thorough research, good expression skills, going through examples online, following the required format, and, most importantly, choosing the best topic. We’ve made that easy with 150 genetics topics you can choose to create exceptional essays in return.
Although this is possible, it can be challenging to source these necessary features alone. That’s where we come in. If you’d like to write an outstanding, best-rated, error-free paper, you can get help with research papers online. We are a group of professional writers with years of writing and research experience. We provide thesis help and full-essay papers with high-quality information for college and university students at a cheap and affordable price.
We write high-quality essays, and we deliver them fast. All you need to do is give us the content of your assignment and other necessary details, including the deadline and your opinions on how you want it executed. You can send a message saying “I need help to write my thesis ” to us online to get an originally written sample at a cheap price.

Make PhD experience your own
Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Main Content
Genetics - research topics.
The following Research Topics are led by experts in their field and contribute to the scientific understanding of genetics. These Research topics are published in the peer-reviewed journal Frontiers in Genetics , as open access articles .

The Role of Growth Factors and Their Receptors in Genetic Disorders
With the advent of next-generation technologies, a large number of genetic diseases have been characterized with the causative genes and gene modifiers. Different developmental anomalies arise as a result of genetic alterations in growth factor genes...

Applications of Mechanistic Modelling for Understanding Human Health and Disease
Mechanistic modeling is a powerful tool for elucidating and probing the behaviors of complex biological systems. By developing mathematical representations based on a deep understanding of biological mechanisms, mechanistic models can provide valuabl...

Understanding Molecular Mechanisms to Facilitate the Development of Biomarkers for Therapeutic Intervention in Gastrointestinal Diseases and Sepsis
This research topic aims to provide an overview of the most recent advancements in understanding molecular mechanisms to develop biomarkers that can be selectively addressed to enhance the diagnosis, prognosis, and therapeutic interventions in gastro...

Integrating Genetics and Proteomics for Drug Discovery
In the past decades, quickly evolving sequencing technology accompanied with big data in computational science has driven a lot of novel medical applications, especially enabling the discovery of associations and causalities. Genome-wide association ...

Advances and Trends in Gene Expression, Regulation, and Phenotypic Variation in Livestock Science: A Comprehensive Review of Methods and Technologies
The introduction of next-generation sequencing (NGS) as well as long-read sequencing technologies has revolutionized the landscape. Over two decades, these technologies have provided high-throughput delivery of genomic data that is more affordable, e...

Nutrients as Regulators of Metabolism: a Genomic Perspective
Our daily diet is more than a collection of carbohydrates, lipids and proteins, minerals and vitamins that provide energy and serve as building blocks of our life, it is also the most dominant environmental signal to which we are exposed from womb to...

Postharvest Ripening, Senescence, and Technology, Volume II
Fresh fruits and vegetables are invaluable for human health and diet, but their quality deteriorates before reaching consumers due to ongoing processes related to ripening and senescence. The field of postharvest biology and technology addresses many...

Challenges and Prospects for Conservation Genetics at XXI Century
Conservation genetics is a fundamental discipline for planning actions for the species and ecosystems management, conservation, and sustainable use. Initially, this work was carried out through collaborative efforts of research groups, sometimes from...

Genetic Testing In Treatment and Management of Alzheimer’s Disease
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder that affects an estimated 50 million people worldwide. This disease is currently the most common cause of dementia in older adults.<br/><br/>Many people wonder if Alzheimer’s diseas...

Recent advances in Genomics and Oncogenomics for Personalized Medicine
Genomic analysis and related molecular analysis technologies undergo rapid advancements, in principle enabling the identification of any genetic alteration potentially implicated in the pathogenesis of diseases and conditions - for germline analyses ...
share this!
April 16, 2024
This article has been reviewed according to Science X's editorial process and policies . Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
Polyploidy in vegetables: Exploring genetics for crop evolution and breeding success
by Chinese Academy of Sciences
A research team has elucidated the role of polyploidy in the evolution and breeding of vegetable crops, leveraging advanced sequencing technologies to dissect the genetic and epigenetic nuances of polyploids. Their findings underline the critical contribution of polyploidy to plant diversity and adaptability, shedding light on "Darwin's abominable mystery" of angiosperm expansion.
Highlighting polyploidy 's potential to enhance crop yields and qualities, the review advocates for its application in vegetable breeding for economic and dietary benefits. Despite the strides in sequencing and multi-omics analysis, challenges persist, including understanding polyploidization's comprehensive impact and leveraging technology for non-model crops.
The study calls for increased research into minor vegetable crops, emphasizing the importance of polyploidy in meeting global food security challenges and promoting a healthy diet amid rising health concerns over high-calorie foods.
Vegetables, crucial for human health and increasingly important economically, are often polyploids, which contribute to their larger organ size and enhanced environmental adaptability. This trait, prevalent in key crops like wheat and cotton, offers significant advantages for breeding, including unique flavors and wider adaptation.
Utilization of advanced sequencing technologies have vastly improved our understanding of vegetable genomics, particularly polyploids, enabling detailed investigations into their evolutionary history and genetic diversity .
Despite these advancements, challenges persist in accurately assembling the complex genomes of polyploids due to their sequence similarity, hindering deeper molecular insights. The current research landscape is poised to further explore the phenotypic advantages and molecular mechanisms of polyploids to untangle the complexities of their genomes, thereby promoting vegetable germplasm innovation and breeding utilization.
A study published in Vegetable Research offers a comprehensive overview of research on vegetable polyploids facilitated by high-throughput sequencing that enhances our grasp of plant evolution and aids in the effective breeding of vegetables through polyploidy.
Genomic sequencing has revolutionized the understanding of vegetable evolution by uncovering ancient whole genome duplication (WGD) events, pivotal for crop domestication and breeding.
Through high-quality genome assemblies, researchers have mapped conserved homologous regions and identified orthologous genes across Cucumis species, revealing the ancient Cucurbit-Common Tetraploidization (CCT) event, which is critical for the divergence of Cucurbitaceae plants.
Despite this, many Cucurbitaceae crops retain genetic information from their diploid progenitors, displaying genetic stability unusual in polyploid evolution. The study of Chinese cabbage and Allium sativum has further highlighted how WGD events contribute to genome expansion and diversification.
Polyploids have complex genomes which make their assembly challenging due to the limitations of sequencing. Recent advances in long-read sequencing have improved the ability to analyze polyploid genomes, revealing structural variants and facilitating the assembly of complex genomes like tetraploid potatoes.
Allopolyploidy arises from the merging of distinct genomes and results in notable changes in gene structure and expression. These changes are gradually stabilized through diploidization, which facilitates species' adaptation to new environments.
This review underscores the potential of polyploidy in enhancing crop diversity and adaptability, essential for addressing global food security. Comparative genomic analyses have shown different fates of polyploid subgenomes, with some becoming dominant, affecting gene expression and agricultural value.
Integrative multi-omics analyses are now enabling efficient crop breeding, offering insights into the complex phenotypes of polyploid vegetables and paving the way for germplasm enhancement through polyploidy.
According to the study's senior researcher, Prof. Xiaqing Yu, "This review summarizes the research progress in vegetable polyploids driven by sequencing technology and the subsequent studies underpinning important traits and genes, which will further promote germplasm innovation and breeding utilization via polyploidy in vegetables."
This work underscores the need for further research, particularly in non-model vegetable crops, to leverage polyploidy for agricultural innovation and to meet the demands of global food security and dietary health.
This calls for more affordable advanced technologies and a greater focus on diversifying vegetable germplasm for a healthier diet, pointing towards a promising direction for polyploidy-based crop breeding aided by artificial intelligence.
Provided by Chinese Academy of Sciences
Explore further
Feedback to editors

Soil bacteria link their life strategies to soil conditions: Study
13 hours ago

Atom-by-atom: Imaging structural transformations in 2D materials
Researchers identify genetic variant that helped shape human skull base evolution
14 hours ago

Two-dimensional nanomaterial sets expansion record

Vibrations of granular materials: Theoretical physicists shed light on an everyday scientific mystery
15 hours ago
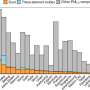

Global study reveals health impacts of airborne trace elements

Researchers find lower grades given to students with surnames that come later in alphabetical order

New model finds previous cell division calculations ignore drivers at the molecular scale

Peptides on interstellar ice: Study finds presence of water molecules not a major obstacle for formation
16 hours ago

Honey bees experience multiple health stressors out in the field
Relevant physicsforums posts, can four legged animals drink from beneath their feet.
Apr 15, 2024
Mold in Plastic Water Bottles? What does it eat?
Apr 14, 2024
Dolphins don't breathe through their esophagus
Is this egg-laying or something else.
Apr 13, 2024
Color Recognition: What we see vs animals with a larger color range
Apr 12, 2024
How to Implement Beamforming in Ultrasound Diffraction Tomography
Apr 10, 2024
More from Biology and Medical
Related Stories


Innovative computational tools provide new insights into the polyploid wheat genome
Feb 27, 2024

Twisted pollen tubes induce infertility in plants with multiple sets of chromosomes
Apr 16, 2024

Polyploidy and long-distance dispersal jointly shape global species diversification in yellowcress, finds study
Dec 22, 2023

Brachypodium model system traces polyploid genome evolution
Aug 26, 2020
Researchers identify role of subgenomes in bamboo evolution
Mar 19, 2024

Unveiling a gap-free genome in rapeseed for enhanced agricultural insight and breeding
Feb 5, 2024
Recommended for you
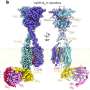
How a calcium-sensing protein multitasks

New class of antimicrobials discovered in soil bacteria
18 hours ago

Neuronal gateway to essential molecules in learning and memory discovered on atomic scale
19 hours ago

How a cyanobacterium manages iron scarcity makes it the most successful photosynthetic organism on Earth
20 hours ago
Let us know if there is a problem with our content
Use this form if you have come across a typo, inaccuracy or would like to send an edit request for the content on this page. For general inquiries, please use our contact form . For general feedback, use the public comments section below (please adhere to guidelines ).
Please select the most appropriate category to facilitate processing of your request
Thank you for taking time to provide your feedback to the editors.
Your feedback is important to us. However, we do not guarantee individual replies due to the high volume of messages.
E-mail the story
Your email address is used only to let the recipient know who sent the email. Neither your address nor the recipient's address will be used for any other purpose. The information you enter will appear in your e-mail message and is not retained by Phys.org in any form.
Newsletter sign up
Get weekly and/or daily updates delivered to your inbox. You can unsubscribe at any time and we'll never share your details to third parties.
More information Privacy policy
Donate and enjoy an ad-free experience
We keep our content available to everyone. Consider supporting Science X's mission by getting a premium account.
E-mail newsletter
- Alzheimer's disease & dementia
- Arthritis & Rheumatism
- Attention deficit disorders
- Autism spectrum disorders
- Biomedical technology
- Diseases, Conditions, Syndromes
- Endocrinology & Metabolism
- Gastroenterology
- Gerontology & Geriatrics
- Health informatics
- Inflammatory disorders
- Medical economics
- Medical research
- Medications
- Neuroscience
- Obstetrics & gynaecology
- Oncology & Cancer
- Ophthalmology
- Overweight & Obesity
- Parkinson's & Movement disorders
- Psychology & Psychiatry
- Radiology & Imaging
- Sleep disorders
- Sports medicine & Kinesiology
- Vaccination
- Breast cancer
- Cardiovascular disease
- Chronic obstructive pulmonary disease
- Colon cancer
- Coronary artery disease
- Heart attack
- Heart disease
- High blood pressure
- Kidney disease
- Lung cancer
- Multiple sclerosis
- Myocardial infarction
- Ovarian cancer
- Post traumatic stress disorder
- Rheumatoid arthritis
- Schizophrenia
- Skin cancer
- Type 2 diabetes
- Full List »
share this!
April 17, 2024
This article has been reviewed according to Science X's editorial process and policies . Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
Electronic health records unlock genetics of tobacco use disorder
by University of California - San Diego
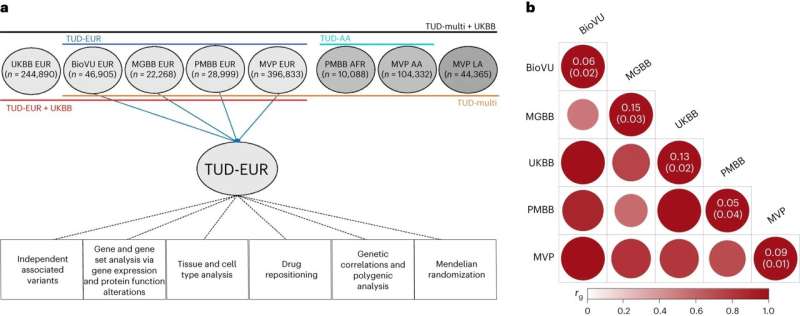
By analyzing electronic health records, researchers at the University of California San Diego School of Medicine have identified hundreds of new genes associated with tobacco use disorder. They also identified hundreds of potential drug candidates that could help treat the disease. The study was published in Nature Human Behavior .
"Tobacco use disorder has an enormous impact on public health," said Sandra Sanchez-Roige, Ph.D., an associate professor in the Department of Psychiatry at UC San Diego School of Medicine. "However, it's challenging to develop new therapeutics for tobacco use disorder because so much of its underlying genetics is poorly understood."
According to the World Health Organization, there are about 1.3 billion tobacco users worldwide, and 80% of these people live in low and middle-income countries. The public health effects of tobacco use extend far beyond those who use it themselves; tobacco kills more than 8 million people each year, and an estimated 1.3 million of these deaths are nonsmokers who were exposed to secondhand smoke .
The official criteria for tobacco use encompass a wide variety of behaviors associated with tobacco use, such as using more tobacco than intended or continuing to use it despite negative consequences. There are known genes associated with nicotine consumption on its own, but these don't tell researchers how nicotine use progresses to tobacco use disorder.
"A fraction of people are able to smoke occasionally without developing an addiction," said Sanchez-Roige. "We want to understand, from a genetic perspective, why occasional tobacco use becomes chronic misuse in some people."
The researchers leveraged large volumes of electronic health data from several health systems in the United States, which was enabled by the PsycheMERGE Network, an international consortium of researchers that aims to synthesize medical records and genomics data to understand better and treat neuropsychiatric illnesses. Sanchez-Roige leads the substance use disorder workgroup within PsycheMERGE.
For the current study, her team used an approach called genome-wide association, which allows researchers to scan the entire genome and look for variations in our genes associated with certain traits, behaviors, or diseases. This is one approach scientists have used to find genes associated with smoking, but this is the first time this approach has been able to reveal genes associated with tobacco use disorder.
In their study of 898,680 individuals, they found 461 candidate risk genes for tobacco use disorder, mostly expressed in the brain. These genes are associated with a myriad of other psychiatric and medical conditions , such as HIV infection, heart disease, and chronic pain. Further, the researchers were able to validate known findings about genes associated with smoking behaviors, which helped validate their approach.
In addition to giving us a more comprehensive view of tobacco use disorder, the researchers were able to use their results to identify hundreds of potential drug candidates that could help doctors treat the disease. However, it will take more research to evaluate these drugs in the lab and the clinic.
In the meantime, the study also supports a growing idea in the field of genetics research: Electronic health records are an underutilized treasure trove of information.
"There's a world of information hidden in medical records, and we accumulate more of them every day as part of routine clinical care," said Sanchez-Roige. "They're also a relatively untapped resource due to how difficult it is to organize and analyze electronic health record data. This study is part of a growing movement to use this constantly expanding source of information to solve complex medical problems."
Explore further
Feedback to editors

Scientists uncover 95 regions of the genome linked to PTSD
2 hours ago

AI tool predicts responses to cancer therapy using information from each cell of the tumor

Health improvements occurred worldwide since 2010 despite COVID-19 pandemic, but progress was uneven: Study
12 hours ago

Study shows heart failure, not stroke is the most common complication of atrial fibrillation
Antipsychotics for dementia linked to more harms than previously acknowledged

Protecting brain cells with cannabinol: Research suggests CBN shows promise for treating neurological disorders
13 hours ago

Calorie restriction study reveals complexities in how diet impacts aging
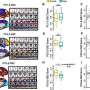

Ketamine produces wide variety of responses in the brain, researchers find
14 hours ago

Emotional radio ads may ease listeners' qualms, boosting support for organ donation

To understand cognition and its dysfunction, neuroscientists must learn its rhythms
Related stories.

Spain bans flavored heated tobacco products
Jan 16, 2024
A deep dive into the genetics of alcohol consumption
Apr 5, 2024

Cigarette smokers more at risk for tobacco dependence than users of smokeless tobacco, multiple tobacco products: Study
Jul 26, 2023

Genomic study links cannabis abuse to multiple health problems
Nov 21, 2023

EU proposes ban on flavored heated tobacco products
Jun 29, 2022
Italy probes BAT, Amazon over heated tobacco ad
Apr 18, 2023
Recommended for you
Shape-shifting cancer cell discovery reveals potential skin cancer drug targets
18 hours ago
Researchers discover how gut muscle can be vital for growth, repair and treatments

Genetic variants found in two types of strabismus, sparking hope for future treatment of eye condition
20 hours ago

New data identifies trends in accidental opioid overdoses in children
17 hours ago
Let us know if there is a problem with our content
Use this form if you have come across a typo, inaccuracy or would like to send an edit request for the content on this page. For general inquiries, please use our contact form . For general feedback, use the public comments section below (please adhere to guidelines ).
Please select the most appropriate category to facilitate processing of your request
Thank you for taking time to provide your feedback to the editors.
Your feedback is important to us. However, we do not guarantee individual replies due to the high volume of messages.
E-mail the story
Your email address is used only to let the recipient know who sent the email. Neither your address nor the recipient's address will be used for any other purpose. The information you enter will appear in your e-mail message and is not retained by Medical Xpress in any form.
Newsletter sign up
Get weekly and/or daily updates delivered to your inbox. You can unsubscribe at any time and we'll never share your details to third parties.
More information Privacy policy
Donate and enjoy an ad-free experience
We keep our content available to everyone. Consider supporting Science X's mission by getting a premium account.
E-mail newsletter
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- HHS Author Manuscripts

Ethical Issues in Genetics and Infectious Diseases Research: An Interdisciplinary Expert Review
Alexis walker.
a Berman Institute of Bioethics, Johns Hopkins University, 1809 Ashland Avenue, Baltimore MD 21205 USA
Vence L. Bonham
b Social and Behavioral Research Branch, National Human Genome Research Institute, 31 Center Drive, Bethesda MD 20894 USA
Angie Boyce
Ellen wright clayton.
c Center for Biomedical Ethics and Society, Vanderbilt University Medical Center, 1211 Medical Center Drive, Nashville TN 37232 USA
Debra Garcia
d International Society for Biological and Environmental Repositories, 750 W Pender St #301, Vancouver BC V6C 1G8 Canada
Stephanie Johnson
e Wellcome Centre for Ethics and the Humanities and Ethox Centre, University of Oxford, Oxford OX1 2JD UK
Oliver Laeyendecker
f Johns Hopkins University School of Medicine, 733 N Broadway, Baltimore MD 21205 USA
g National Institute of Allergy and Infectious Diseases, National Institutes of Health, 5601 Fishers Ln, Bethesda, MD 20892 USA
Michelle Lewis
Joseph b. margolick.
h Johns Hopkins University Bloomberg School of Public Health, 615 N Wolfe St, Baltimore MD 21205 USA
Debra Mathews
Michael j. parker, paul spicer.
I Department of Anthropology and the Center for Applied Social Research, University of Oklahoma, 455 W Lindsey St, Norman OK 73069 USA
Chloe L. Thio
Gail geller, jeffrey kahn.
Author Contributions
Research in genetics and infectious diseases (ID) presents novel configurations of ethical, legal, and social issues (ELSIs) related to the intersection of genetics with public health regulations and the control of transmissible diseases. Such research includes work both in pathogen genetics and on the ways that human genetics affect responses to ID. This paper identifies and systematizes the unique issues at this intersection, based on an interdisciplinary expert review.
Basic Procedures.
This paper presents results of a formal issue-spotting exercise among twenty experts in public health, law and genomics, biobanking, genetic epidemiology, ID medicine and public health, philosophy, ethics and ID, ethics and genomics, and law and ID. The focus of the exercise was on the collection, storage, and sharing of genetic information relating to ID.
Main Findings.
The issue-spotting exercise highlighted the following ELSIs: risks in reporting to government authorities, return of individual research results, and resource allocation – each taking on specific configurations based on the balance between public health and individual privacy/protection.
Principal Conclusions.
The public health implications of interactions between genomics and ID frame considerations for equity and justice. In the context of the COVID-19 pandemic, these issues are especially pressing.
Introduction
Much has been learned in recent years about how pathogen and human genetics affect the incidence and outcomes of infectious diseases [ 1 ]. And while the COVID-19 pandemic underlines the importance of this knowledge, it also highlights pressing ethical, legal, and social issues (ELSIs) in genomics and infectious disease. These include data privacy, allocation of medical resources, and over-emphasis on genetic drivers of inequity as opposed to social factors – all longstanding issues that take on novel permutations at the intersection of genomics and infectious diseases research. While this area of research and practice is subject to the extensive ELSIs that have been described in genomics and non-communicable disease [ 2 – 4 ], new formulations occur as disparate types of information from the fields of genetics and of infectious disease are brought together at unprecedented extents and levels of detail, which potentially multiplies risks [ 5 – 7 ].
This paper presents results of an issue-spotting exercise conducted by experts in the ethics, law, and science of genetics and infectious diseases (ID). The exercise focused on the collection, storage and sharing of genetic data relating to ID, highlighting ELSIs that differ in important ways from issues in genetics and non-transmissible disease. While this exercise was conducted prior to the emergence of COVID-19, the issues raised have become all the more urgent in the context of the current pandemic.
Materials and Methods
The primary method of this study is a systematic issue-spotting exercise with twenty expert participants, conducted according to validated methods in law and bioethics [ 8 ]. This exercise was organized as part of a [BLIND] grant funded at [BLIND] by the National Human Genome Research Institute (NHGRI). Issue-spotting is a commonly used method for identifying a range of ethical issues that might arise in the near to mid-term future, based on anticipated developments in law and science. In the current study, the issue-spotting exercise involved the exploration of possible ELSIs by a group of 20 experts from the following fields: public health, law and genomics, biobanking, genetic epidemiology, ID medicine and public health, philosophy, ethics and ID, ethics and genomics, and law and ID. Members of the group represented a diversity of expertise relevant to our focus and are based at universities and organizations across the United States and the United Kingdom.
The issue-spotting was conducted through a series of three meetings in Spring 2019: a two-day in-person conference held at [BLIND], and two half-day WebEx conferences. Before meetings, the group reviewed relevant literature on ethics in genomics and ID to inform development of a series of scenarios, which the group then used to help identify areas of ethical concern. This effort included imagining possible developments based on current research, and the ELSI issues such developments would present [ 9 ]. Analysis was collaborative among the participant group; all participants shared perspectives on possible and likely directions for genomics and ID and the ELSIs they present, discussing any disagreements within the group. After broad and extensive live discussion (more than ten hours, spread over two months), the group collaboratively determined the issues that had been most discussed, and agreed on a baseline set of consensus concerns, described below.
Authors of this paper are a subset of the full group involved in the issue-spotting. The issues identified are largely focused on ethical considerations, but some legal elements are also noted.
As genomics becomes increasingly important to ID research–for example, revealing individuals at lower or greater genetic risk of being infected with specific pathogens or more or less likely to heal quickly from infections [ 10 ] – ID research has had to grapple with ELSIs common to the broader field of genomics, such as genetically-based group harm or stigma [ 11 ], and ownership of genetic data or samples [ 12 ]. However, the close relationship of ID with public health interventions has led to a different configuration of the privacy and justice concerns that pertain to other areas of genetics research. The Working Group identified three primary areas of ELSI concern of this type: reporting results to authorities, return of individual research results, and allocation of resources for clinical care.
1. Reporting Results to Authorities
In order to track disease trends and outbreaks, public health authorities require that cases of ID of public health concern be reported to local, regional, and national agencies. The US Centers for Disease Control and Prevention, for example, require reporting of over sixty IDs by medical professionals and laboratories. The reportability of ID presents unique concerns when ID and genetics research come together. For example, if human genetic variations associated with increased risk of transmitting an infection were to be identified, public health authorities could potentially require that such genetic information be reported for cases of that infection. As seen for COVID-19, there has been substantial interest in early identification of “superspreaders” based on both behavioral and viral genetic factors [ 13 ]. Host genetic information could present a means of pre-emptive identification of individuals or groups at high risk of spreading a virus in an attempt to control that infection [ 14 ], with a host of attending concerns. Such information could be used to limit the ability of some individuals to work outside their homes. This would present pressing questions regarding thresholds (e.g., what level of evidence and increased risk could justify such decisions?) and justice (e.g., what financial support should be provided to people who are not able to work, and who should provide it?).
In addition to human genetic information, public health authorities might also be interested in pathogen genetic information. Paths of disease transmission identified by tracing evolutionary relationships among pathogen samples (phylogenetics) are used by public health authorities to track disease [ 15 ]. Public health agencies have long used contact tracing, based on the names of contacts provided by an affected person, to follow up with potentially infected individuals and control disease transmission. For HIV, pathogen sequence data are assessed for the formation of viral clusters, which if found among newly diagnosed individuals, can trigger augmented partner tracing [ 16 ]. The significance of pathogen genetics in tracing disease spread has led some public health agencies to collect not only information on disease cases but also the genetic sequence of the case’s pathogen and associated meta-data – as seen in public health surveillance of SARS-CoV-2 in some countries [ 13 ]. Although such meta-data is generally de-identified, in some cases information about an individual human host can be derived from the pathogen sequence itself.
Even though sharing genetic information relating to ID could reduce risk to public health, it also can raise concerns about privacy. Mandatory reporting of human and pathogen genetic information on ID would limit the control research participants have over data about themselves; this is different from traditional contact tracing, in which individuals ultimately retain power over the information they share. Participants in long-term HIV cohort studies have expressed confidence in the clinical researchers they interact with directly, but also substantial concern about use of their data by government entities [ 6 ], highlighting the potential importance of control over information shared.
When pathogen and human genetic information are combined, risks to privacy can increase. Information about pathogen spread is often a powerful indication of social interactions and behavior -- information that can potentially be used against people. For example, this information may place people in legal jeopardy when shared with government agencies, as it is illegal in some jurisdictions to expose others to an infectious agent, and genetic information can, in some contexts, enable confirmation of a person who has transmitted a pathogen [ 17 ].
While public health authorities in the United States and elsewhere have done an admirable job of protecting infectious disease information, the possibility that genetic and ID information could be accessed by other government agencies is a serious risk, especially for members of groups who have been, and continue to be, disproportionately targeted by police, who stand to lose public benefits, and/or who are undocumented. These issues arise for both pathogen and human genetic information relating to ID, especially when these kinds of information can be cross-referenced and thus can reveal more about individuals [ 19 ]. The protection of sensitive information by public health authorities has at times depended on the actions of individuals who stood up to demands for access by law enforcement [ 6 ]. This is a thin protection.
2. Return of Individual Research Results
Today there is a widespread expectation that researchers will share with research participants aggregate data from the studies in which these participants take part; in addition, there has been growing pressure for expanded return of individual research results [ 20 ]. The latter has been based on a) the desire participants have expressed to receive this information, b) broadening understanding of rights to health information, and c) a desire to balance the increased privacy risks in genetics research with potential benefits. For example, some participants in a longitudinal HIV cohort wanted to know whether they carried a mutation in the CCR5 gene that could affect acquisition and progression of HIV infection. The researchers, after consulting with their IRB, made these results available to the participants as a service to them, even though this information would not affect their health care or their benefit from practicing safe sex.
IRBs and researchers have expressed concern that sharing individual research results about genetic variants that are protective against a specific ID could lead people to engage in riskier behavior. This could then contribute to increased transmission of that or other diseases [ 8 ]; similar concerns were raised in connection with use of pre-exposure prophylaxis (PrEP) for HIV infection. However, there is currently no evidence of such increased risk-taking based on results relating to the genetics of ID – either host or pathogen.
Participants in genetics ID research report great interest in their individual results, not only for their own knowledge, but to better protect those around them [ 6 ]. Researchers should weigh sincere respect for participant views against the effects of returning individual results, including possible effects on cost of the research enterprise and health care, which would be increased by expenses to adequately confirm tests, track individual participants, and return results [ 21 ]. These questions require more systematic discussion within research communities to ensure that researchers are making well-considered decisions in managing return of individual results.
3. Allocation of Resources
Genetic information regarding people’s susceptibility to or transmission of specific IDs may also affect the allocation of scarce resources. In the case of hepatitis C, a few researchers have suggested that patients with higher genetic likelihood of response to interferon-based therapies (which are generally poorly tolerated) or a higher likelihood of spontaneous viral clearance could be deprioritized from treatment with highly effective, but very costly, medications that can cure hepatitis C [ 6 ].
In the future some stakeholders might become more interested in such “precision rationing” approaches for a variety of conditions and resources [ 6 ]. For example, patients’ genetic predisposition to influenza (or COVID-19) progression and spread could be used to allocate isolation beds [ 19 ]. But how might such approaches contribute to health inequities? This calculus is made more complex by the question of whom the intervention is protecting – the patient and/or others? Concerns for efficient resource allocation should not inhibit prioritization of justice and the importance of countering current health inequities; funding must be made available to support health justice for all.
Here again, human genetics of ID presents public health issues that are not commonly considered in human genetics of non-communicable disease, which has been the focus of past ELSI research. Further anticipatory ELSI research is necessary to investigate where, and how, genetics-based rationing might be applied in both the short and longer term, to ensure that resource allocation achieves distributive justice.
Discussion/Conclusion
Burgeoning research on genetics of human ID and of human pathogens presents novel configurations of ethical, legal, and social issues in genetics. Specifically, public health concerns lead to the possibility that genetic information related to ID could be used in attempts to control transmission, involving relationships of authority not engaged in the management of non-communicable disease. The Covid-19 pandemic has highlighted vast inequalities in the management of infectious disease and in the social context that underlies human health. Researchers have the ability and responsibility to confront additional issues in the research process at the intersection of genomics and ID research – regarding, for example, reporting data to authorities, return of individual research results, and resource allocation. These areas require norms and policies to protect privacy and advance equity beyond current protections. This necessitates sustained research and public discussion to protect the most marginalized and to incorporate both expert and lay perspectives, especially those from people who have borne, and still bear, the negative effects of discrimination -- based on gender, race, ethnicity, immigration status, disability, age, sexual orientation, or income. Coupling such research and discussion with action is crucial for preventing adverse and undesired effects of research, including impaired privacy and legal safety of research participants, loss of trust in the research enterprise, harm and stigmatization of specific groups of people, and exacerbation of or failure to improve health inequities.
Acknowledgements
This paper is submitted on behalf of the Working Group on ELSI, Biobanks, Infectious Diseases, and Marginalized Groups, with important contributions by Seema Shah, Margaret Battin, Leslie Meltzer Henry, and Shruti Mehta – in addition to the named authors.
Funded as part of the NHGRI CEER II RM1HG009038 - Ethical, Legal & Social Implications of "Precision Medicine" in Infectious Disease, whose primary investigators are authors on this paper (Geller and Kahn). The funder has had no role in the analysis or in the preparation of this manuscript.
Conflict of Interest
The authors have no conflicts of interest to declare.
Statement of Ethics
This paper is based on methods for ethics analysis and does not constitute research with human subjects.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
Genomics articles from across Nature Portfolio
Genomics is the study of the full genetic complement of an organism (the genome). It employs recombinant DNA, DNA sequencing methods, and bioinformatics to sequence, assemble, and analyse the structure and function of genomes.
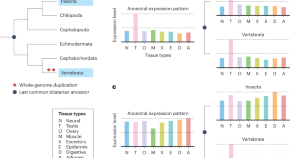
Tissue specificity follows gene duplication
A comparative transcriptomic analysis of eight tissue types in twenty bilaterian species reveals the long-lasting effects of genome duplication on the evolution of novel tissue-specific gene-expression patterns.
- Anamaria Necsulea
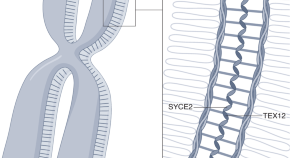
A rare genetic variant biases maternal meiotic recombination toward risk of pregnancy loss
Pregnancy loss is common in humans, but maternal genetic factors modulating its incidence are largely unknown. In a meta-analysis of genome-wide association studies, researchers identified a genetic variant that seems to increase risk of pregnancy loss by dysregulating meiotic recombination between homologous chromosomes during egg formation.
- Sara A. Carioscia
- Rajiv C. McCoy
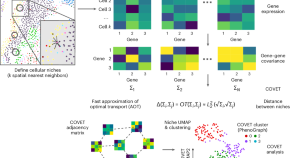

Capturing and modeling cellular niches from dissociated single-cell and spatial data
Cells interact with their local environment to enact global tissue function. By harnessing gene–gene covariation in cellular neighborhoods from spatial transcriptomics data, the covariance environment (COVET) niche representation and the environmental variational inference (ENVI) data integration method model phenotype–microenvironment interplay and reconstruct the spatial context of dissociated single-cell RNA sequencing datasets.
Related Subjects
- Comparative genomics
- Conservation genomics
- Epigenomics
- Genome assembly algorithms
- Genome evolution
- Medical genomics
- Metagenomics
- Mobile elements
- Nutrigenomics
- Personalized medicine
- Pharmacogenomics
- Phylogenomics
- Transcriptomics
Latest Research and Reviews

Genetic modifiers of rare variants in monogenic developmental disorder loci
Analysis of genetic modifiers of 599 developmental disorder genes in the UK Biobank found that rare variant burden within this set, as well as the common polygenic background, can alter the expressivity of cognitive and socioeconomic traits in an additive manner.
- Rebecca Kingdom
- Robin N. Beaumont
- Caroline F. Wright
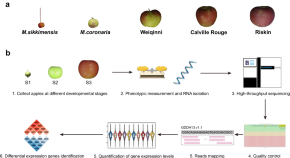
Phenomics and transcriptomic profiling of fruit development in distinct apple varieties
- Weihan Zhang
- Yuepeng Han
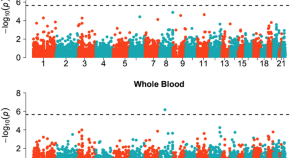
Investigating the role of common cis -regulatory variants in modifying penetrance of putatively damaging, inherited variants in severe neurodevelopmental disorders
- Emilie M. Wigdor
- Kaitlin E. Samocha
- Hilary C. Martin
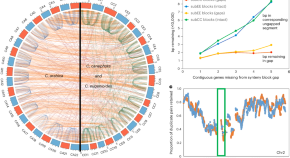
The genome and population genomics of allopolyploid Coffea arabica reveal the diversification history of modern coffee cultivars
Chromosome-level genome assemblies of allotetraploid Coffea arabica and representatives of its diploid progenitors, Coffea eugenioides and Coffea canephora , provide insights into Arabica’s diversification history.
- Jarkko Salojärvi
- Aditi Rambani
- Patrick Descombes
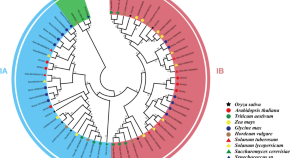
Functional differentiation and genetic diversity of rice cation exchanger (CAX) genes and their potential use in rice improvement
- Shangshu Lian
- Yanjun Chen
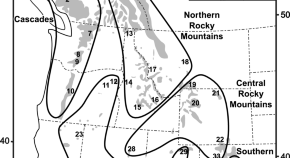
Putative climate adaptation in American pikas ( Ochotona princeps ) is associated with copy number variation across environmental gradients
- Bryson M. F. Sjodin
- Danielle A. Schmidt
- Michael A. Russello
News and Comment
Quantitative proteomics of protein–nucleosome interactions.
- Michael Fletcher
Implementing polygenic risk scores in the clinic
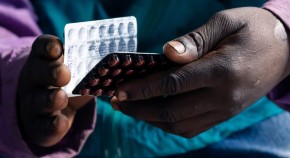
AI can help to tailor drugs for Africa — but Africans should lead the way
Computational models that require very little data could transform biomedical and drug development research in Africa, as long as infrastructure, trained staff and secure databases are available.
- Gemma Turon
- Mathew Njoroge
- Kelly Chibale
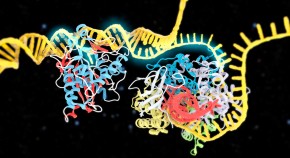
Advanced CRISPR system fixes a deadly mutation in cells
Applying a ‘base editor’ allows cells to crank out increased levels of a vital metabolic enzyme.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Thanks to a genetic breakthrough, a rare rhino species may be rescued from extinction
New research from the san diego zoo using just 12 frozen cells suggests we can protect the northern white rhino, by matthew rozsa.
As humans continue to encroach on our planet, we are driving a mass extinction that some experts call a " biological holocaust ." Since more and more species are dying, it creates an increasing number of genetic bottlenecks, which make animal and plant survival even more difficult.
Take for example the white rhinoceros ( Ceratotherium simum ), which can be divided into two sub-species that are genetically very similar — but one is relatively thriving while the other is on the brink of extinction.
Scientists using state-of-the-art genetics technology hope to change that — and their recent research published in the journal Evolutionary Applications suggests they might be able to pull it off.
"Doing our best to preserve [rhinos] is really our moral obligation."
Such a feat would not be unprecedented. The southern white rhinoceroses, which is currently the most abundant rhinoceros species in the world, had been culled down to a population of merely 50 to 200 individuals in the early 20th century. Thanks to rigorous conservation efforts, however, the southern white rhinoceros population had rebounded to roughly 20,000 individuals by 2014 (a surge in poaching around that time has since reduced their population to roughly 18,000).
The southern white rhinoceros' cousin, however, faces a much more dire situation. At the time of this writing, there are only two females from the northern white rhinoceros species that are still alive. Even if there was a male around, it would not matter, since both females are past the age when they can carry a fetus to term. Poaching, poorly managed land use and other human activities have taken a massive toll.
While a few decades ago this would have entirely doomed the species, cutting edge advances in genetics technology may offer them salvation. Dr. Aryn P. Wilder — a conservation scientist at the San Diego Zoo Wildlife Alliance — decided to study genetic samples from 12 northern white rhinoceroses that had cytogenetically frozen at the San Diego Zoo. Much to her delight, Wilder found that those dozen samples contain enough genetic diversity that one could resurrect them from functional extinction.
Indeed, not only is there enough diversity to allow rhinoceroses to be reproduced through cloning, but the samples from northern white rhinoceroses are actually more diverse than those of the southern white rhinoceroses. This means that if scientists are able to bring them back, they will be less likely to encounter a genetic bottleneck, in which individual animals are born unhealthy because their parents are too closely related to each other.
Salon spoke with Wilder about this uplifting news, as well as the practical steps that need to be taken next to save northern white rhinoceroses.
This interview has been lightly edited for length and clarity.
Can you explain how your technology was able to determine that there is enough genetic diversity within these 12 samples to avoid a genetic bottleneck?
We sequenced the genomes of individuals from both species, northern white rhinos and southern white rhinos. When you sequence the genome, you can actually measure the amount of genetic diversity in each of those genomes. And we know that the southern white rhino was able to recover without too much inbreeding. So we used southern white rhinos as a benchmark or metric of a healthy enough population. And so then we asked, "Well, do the cells that we have banked in the frozen zoo, do they have enough genetic diversity to recover in a similar way?"
What we found was that, yes, the northern white rhinos actually have more genetic diversity in their genomes than the southern white rhinos, so we know then that they at least have adequate levels of genetic diversity. The other thing that we wanted to look at was harmful mutations in the genome. So we can actually look at genetic variants in the genome or mutations in the genome, and predict how harmful they might be. If those mutations are in a gene that encodes a protein, we can predict what the protein will look like, and we know that if a mutation causes a change in that protein, it's more likely to be harmful.
We need your help to stay independent
We can also look across lots of mammals in other mammal species. If we find that that mutation is very rare or doesn't exist in any other mammal species, and every other mammal species has the same genetic variant, then we would infer that that mutation is actually really important, or that that position is really important in any change to that position and is likely harmful.
Then we counted up all of the mutations in the northern white rhino genome that were harmful and did the same in the southern white rhino, and then modeled over time what those mutations would do in a restored population and whether those harmful mutations would accumulate and cause fitness declines that made the northern white rhino's fitness lower than the southern white rhino.
Then in order to predict what those mutations would do when a northern white rhino population is restored from banked cells in the frozen zoo, we used genomic simulations and we said, "Okay, well if we were to take eight of those individuals from the frozen zoo, clone them and start a population of over 10 generations, what would that look like? What would fitness look like in generation 1, 2, 3 and all the way through generation 10?"
And then we also modeled taking those same eight individuals, starting the population in generation zero, and then every generation after that we introduced one new cell line into — or one new cloned individual back into — the population. Basically modeling this regenerative source of genetic diversity, or this bank of genetic diversity that we have in the frozen zoo, and we found that the populations that had founders reintroduced every generation, they did much better. They didn't suffer any fitness declines like the ones that were just founded once in generation zero and then allowed to to from there over the next 10 generations.
Want more health and science stories in your inbox? Subscribe to Salon's weekly newsletter Lab Notes .
"We still need to to use all of the traditional conservation methods that we've always used."
What are the next steps now that this technology has demonstrated to work?
What these models have shown is that the source of genetic diversity that we have in these cells is enough to restore a healthy population. But in order to restore a healthy population, we need to be able to use these cell lines and actually clone northern white rhino embryos, or create northern white rhino sperm and eggs that then we can use for in vitro fertilization to make an embryo from there.
We need to implant the embryo into a southern white rhino surrogate mother, and she needs to carry her baby to term, and then critically we need to have a habitat that we can release these rhinos into in the wild. They need to have all of the protections that should have been given to them before their population was reduced to just two females. They need to have protection from poaching. They need to have adequate space and healthy habitats for them to live in.
Do you have any personal stories of interactions you've had with rhinos through your research?
Well, we do have a herd of southern white rhinos here. I've actually never interacted with the northern white rhino because by the time I started at the zoo, there were only three left. There was a male, but he passed away a few years ago. Now it's just the two left. But from what I've seen of the southern white rhinos, the closely related subspecies, they're a very gentle and sweet species. That's not to say that that in the wild they'll be gentle and sweet with you, but I've seen the moms with their babies, and the babies wallowing in the mud, and they're really a unique species — doing our best to preserve them is really our moral obligation.
I want to emphasize again that these new sorts of cellular technologies are only one tool in the toolbox. We still need to to use all of the traditional conservation methods that we've always used. Like I said before, we still need to protect these species and their habitats. We can't just expect that these methods are sufficient to save a species and to end the extinction crisis. This is just one tool in the toolbox, and the reason we have this tool for this species is because we thought ahead to bank these cells. There are increasing efforts to create these biobanks of living cells so that we can have this genetic material for the future. I think that banking species, even before banking cells from species, even before they suffer these really severe declines, is going to be a really critical resource for the future.
about conservation
- Losing hippos could throw entire ecosystems out of balance. Here's how we can still save them
- Western wolves have made a comeback — now Western states want to kill them
- Monkeys are marvelous, but face a bleak future thanks to other primates: Humans
Matthew Rozsa is a staff writer at Salon. He received a Master's Degree in History from Rutgers-Newark in 2012 and was awarded a science journalism fellowship from the Metcalf Institute in 2022.
Related Topics ------------------------------------------
Related articles.
- MyU : For Students, Faculty, and Staff
Fall 2024 CSCI Special Topics Courses
Cloud computing.
Meeting Time: 09:45 AM‑11:00 AM TTh Instructor: Ali Anwar Course Description: Cloud computing serves many large-scale applications ranging from search engines like Google to social networking websites like Facebook to online stores like Amazon. More recently, cloud computing has emerged as an essential technology to enable emerging fields such as Artificial Intelligence (AI), the Internet of Things (IoT), and Machine Learning. The exponential growth of data availability and demands for security and speed has made the cloud computing paradigm necessary for reliable, financially economical, and scalable computation. The dynamicity and flexibility of Cloud computing have opened up many new forms of deploying applications on infrastructure that cloud service providers offer, such as renting of computation resources and serverless computing. This course will cover the fundamentals of cloud services management and cloud software development, including but not limited to design patterns, application programming interfaces, and underlying middleware technologies. More specifically, we will cover the topics of cloud computing service models, data centers resource management, task scheduling, resource virtualization, SLAs, cloud security, software defined networks and storage, cloud storage, and programming models. We will also discuss data center design and management strategies, which enable the economic and technological benefits of cloud computing. Lastly, we will study cloud storage concepts like data distribution, durability, consistency, and redundancy. Registration Prerequisites: CS upper div, CompE upper div., EE upper div., EE grad, ITI upper div., Univ. honors student, or dept. permission; no cr for grads in CSci. Complete the following Google form to request a permission number from the instructor ( https://forms.gle/6BvbUwEkBK41tPJ17 ).
CSCI 5980/8980
Machine learning for healthcare: concepts and applications.
Meeting Time: 11:15 AM‑12:30 PM TTh Instructor: Yogatheesan Varatharajah Course Description: Machine Learning is transforming healthcare. This course will introduce students to a range of healthcare problems that can be tackled using machine learning, different health data modalities, relevant machine learning paradigms, and the unique challenges presented by healthcare applications. Applications we will cover include risk stratification, disease progression modeling, precision medicine, diagnosis, prognosis, subtype discovery, and improving clinical workflows. We will also cover research topics such as explainability, causality, trust, robustness, and fairness.
Registration Prerequisites: CSCI 5521 or equivalent. Complete the following Google form to request a permission number from the instructor ( https://forms.gle/z8X9pVZfCWMpQQ6o6 ).
Visualization with AI
Meeting Time: 04:00 PM‑05:15 PM TTh Instructor: Qianwen Wang Course Description: This course aims to investigate how visualization techniques and AI technologies work together to enhance understanding, insights, or outcomes.
This is a seminar style course consisting of lectures, paper presentation, and interactive discussion of the selected papers. Students will also work on a group project where they propose a research idea, survey related studies, and present initial results.
This course will cover the application of visualization to better understand AI models and data, and the use of AI to improve visualization processes. Readings for the course cover papers from the top venues of AI, Visualization, and HCI, topics including AI explainability, reliability, and Human-AI collaboration. This course is designed for PhD students, Masters students, and advanced undergraduates who want to dig into research.
Registration Prerequisites: Complete the following Google form to request a permission number from the instructor ( https://forms.gle/YTF5EZFUbQRJhHBYA ). Although the class is primarily intended for PhD students, motivated juniors/seniors and MS students who are interested in this topic are welcome to apply, ensuring they detail their qualifications for the course.
Visualizations for Intelligent AR Systems
Meeting Time: 04:00 PM‑05:15 PM MW Instructor: Zhu-Tian Chen Course Description: This course aims to explore the role of Data Visualization as a pivotal interface for enhancing human-data and human-AI interactions within Augmented Reality (AR) systems, thereby transforming a broad spectrum of activities in both professional and daily contexts. Structured as a seminar, the course consists of two main components: the theoretical and conceptual foundations delivered through lectures, paper readings, and discussions; and the hands-on experience gained through small assignments and group projects. This class is designed to be highly interactive, and AR devices will be provided to facilitate hands-on learning. Participants will have the opportunity to experience AR systems, develop cutting-edge AR interfaces, explore AI integration, and apply human-centric design principles. The course is designed to advance students' technical skills in AR and AI, as well as their understanding of how these technologies can be leveraged to enrich human experiences across various domains. Students will be encouraged to create innovative projects with the potential for submission to research conferences.
Registration Prerequisites: Complete the following Google form to request a permission number from the instructor ( https://forms.gle/Y81FGaJivoqMQYtq5 ). Students are expected to have a solid foundation in either data visualization, computer graphics, computer vision, or HCI. Having expertise in all would be perfect! However, a robust interest and eagerness to delve into these subjects can be equally valuable, even though it means you need to learn some basic concepts independently.
Sustainable Computing: A Systems View
Meeting Time: 09:45 AM‑11:00 AM Instructor: Abhishek Chandra Course Description: In recent years, there has been a dramatic increase in the pervasiveness, scale, and distribution of computing infrastructure: ranging from cloud, HPC systems, and data centers to edge computing and pervasive computing in the form of micro-data centers, mobile phones, sensors, and IoT devices embedded in the environment around us. The growing amount of computing, storage, and networking demand leads to increased energy usage, carbon emissions, and natural resource consumption. To reduce their environmental impact, there is a growing need to make computing systems sustainable. In this course, we will examine sustainable computing from a systems perspective. We will examine a number of questions: • How can we design and build sustainable computing systems? • How can we manage resources efficiently? • What system software and algorithms can reduce computational needs? Topics of interest would include: • Sustainable system design and architectures • Sustainability-aware systems software and management • Sustainability in large-scale distributed computing (clouds, data centers, HPC) • Sustainability in dispersed computing (edge, mobile computing, sensors/IoT)
Registration Prerequisites: This course is targeted towards students with a strong interest in computer systems (Operating Systems, Distributed Systems, Networking, Databases, etc.). Background in Operating Systems (Equivalent of CSCI 5103) and basic understanding of Computer Networking (Equivalent of CSCI 4211) is required.
- Future undergraduate students
- Future transfer students
- Future graduate students
- Future international students
- Diversity and Inclusion Opportunities
- Learn abroad
- Living Learning Communities
- Mentor programs
- Programs for women
- Student groups
- Visit, Apply & Next Steps
- Information for current students
- Departments and majors overview
- Departments
- Undergraduate majors
- Graduate programs
- Integrated Degree Programs
- Additional degree-granting programs
- Online learning
- Academic Advising overview
- Academic Advising FAQ
- Academic Advising Blog
- Appointments and drop-ins
- Academic support
- Commencement
- Four-year plans
- Honors advising
- Policies, procedures, and forms
- Career Services overview
- Resumes and cover letters
- Jobs and internships
- Interviews and job offers
- CSE Career Fair
- Major and career exploration
- Graduate school
- Collegiate Life overview
- Scholarships
- Diversity & Inclusivity Alliance
- Anderson Student Innovation Labs
- Information for alumni
- Get engaged with CSE
- Upcoming events
- CSE Alumni Society Board
- Alumni volunteer interest form
- Golden Medallion Society Reunion
- 50-Year Reunion
- Alumni honors and awards
- Outstanding Achievement
- Alumni Service
- Distinguished Leadership
- Honorary Doctorate Degrees
- Nobel Laureates
- Alumni resources
- Alumni career resources
- Alumni news outlets
- CSE branded clothing
- International alumni resources
- Inventing Tomorrow magazine
- Update your info
- CSE giving overview
- Why give to CSE?
- College priorities
- Give online now
- External relations
- Giving priorities
- Donor stories
- Impact of giving
- Ways to give to CSE
- Matching gifts
- CSE directories
- Invest in your company and the future
- Recruit our students
- Connect with researchers
- K-12 initiatives
- Diversity initiatives
- Research news
- Give to CSE
- CSE priorities
- Corporate relations
- Information for faculty and staff
- Administrative offices overview
- Office of the Dean
- Academic affairs
- Finance and Operations
- Communications
- Human resources
- Undergraduate programs and student services
- CSE Committees
- CSE policies overview
- Academic policies
- Faculty hiring and tenure policies
- Finance policies and information
- Graduate education policies
- Human resources policies
- Research policies
- Research overview
- Research centers and facilities
- Research proposal submission process
- Research safety
- Award-winning CSE faculty
- National academies
- University awards
- Honorary professorships
- Collegiate awards
- Other CSE honors and awards
- Staff awards
- Performance Management Process
- Work. With Flexibility in CSE
- K-12 outreach overview
- Summer camps
- Outreach events
- Enrichment programs
- Field trips and tours
- CSE K-12 Virtual Classroom Resources
- Educator development
- Sponsor an event
- See us on facebook
- See us on twitter
- See us on youtube
- See us on linkedin
- See us on instagram
Pediatric rheumatologist Elizabeth Mellins dies at 72
Mellins, who studied autoimmune disease and co-founded a large pediatric rheumatology research network, was a tireless mentor and advocate for her field.
April 12, 2024 - By Erin Digitale

Betsy Mellins “wanted other people to care about the science as much as she cared, because it was science that would lead to better treatments for children,” Jennifer Frankovich said. Stanford Medicine
Elizabeth “Betsy” Mellins, MD, professor of pediatrics at the Stanford School of Medicine, died March 24. She was 72.
Mellins was a pediatric rheumatologist and immunologist whose laboratory made important discoveries about childhood inflammatory diseases. She was also a dedicated advocate for her specialty, a founder of one of its largest research networks, and a devoted and tenacious mentor to young scientists and physicians.
“Dr. Mellins was a consummate physician-scientist whose research acumen was driven by her deep compassion for children living with juvenile arthritis and other autoimmune diseases,” said Lloyd Minor , MD, dean of the Stanford School of Medicine and vice president for medical affairs at Stanford University. “Not only did she make important discoveries in her own lab, she worked tirelessly to expand research opportunities for everyone in her field, mentoring so many physicians to the ultimate benefit of young patients who can now access safer and more effective treatments for challenging chronic illnesses.”
Mellins tackled several difficult problems in immunology, focusing her research on a form of childhood arthritis, especially how immune markers known as MHC class II molecules function as inherited risk factors. Her recent work helped physicians understand an emerging, life-threatening lung complication of systemic juvenile idiopathic arthritis, or Still disease. Although it is not fully understood, scientists hypothesize that some newer drugs cause an immune-system imbalance in certain patients that leads to this complication. Mellins also gathered evidence for the autoimmune underpinnings of a severe pediatric psychiatric condition called pediatric acute-onset neuropsychiatric syndrome, or PANS. When her findings ran counter to perceived wisdom, she was adept at overcoming others’ skepticism.
“She was brilliant: She had the gift of not only knowing the science, but how to share it with the rest of the world,” said Jennifer Frankovich , MD, clinical professor of pediatrics and director of the Stanford PANS research program and the Immune Behavioral Health Clinic at Lucile Packard Children’s Hospital Stanford. “She wanted other people to care about the science as much as she cared, because it was science that would lead to better treatments for children.”
Having entered her profession at a time when female physician-scientists were rare, she worked hard to expand opportunities for young researchers, especially women.
“She loved finding and nurturing bright, interested young talent: undergrads, PhD students, postdocs, residents and fellows,” said Vivian Saper, MD, adjunct clinical professor of pediatrics. “Anybody who had gone through Betsy’s lab had a friend and mentor for life.”
‘Most likely to succeed’
Mellins was born Dec. 16, 1951, in Minneapolis, and grew up in Manhasset, New York, as the eldest child of an academically oriented family. Her father, Harry Mellins, MD, was a radiologist on the faculty of Harvard Medical School, and her mother, Judith Weiss Mellins, held a master’s degree in economics from Radcliffe College, which would later become part of Harvard. Mellins had two younger brothers, Bill and Tom.
After completing high school in 1969, Mellins earned an undergraduate degree from Cornell University in political science. While at Cornell, she enrolled for a summer quarter at Stanford and fell in love with it. She thought about becoming a teacher, but changed course.
“She had been voted ‘most likely to succeed’ in high school, and the best possible career path for a brilliant woman in 1969 was schoolteacher,” said her daughter, Lisa Mendelman.
Instead, Mellins did a post-baccalaureate year at MIT and went to Harvard Medical School, graduating in 1978. She then completed a pediatric residency at the University of Colorado Health Sciences Center in Denver, and a final year of residency and clinical fellowship in pediatric rheumatology at the University of Washington in Seattle, where she went on to a postdoctoral research fellowship in developmental biology.
While in medical school, Mellins spent a summer in the Sonoran Desert on the Tohono Oʼodham reservation in Sells, Arizona, gaining clinical experience. There, she met Paul Mendelman, MD, a physician in the U. S. Public Health Service. They were together for 48 years, marrying in 1980.
“Years later, she would say that if there had been mentors available, she would have been mentored to get a PhD, not an MD, based on her interests,” Lisa Mendelman said. “She was striking out into such unprecedented territory. It explains her lifelong commitment to scientific mentorship, to giving other generations experiences she did not have.”
Research focus
After completing her training, Mellins held faculty positions at the University of Washington and the University of Pennsylvania before being recruited to Stanford Medicine as an associate professor of pediatrics in 1996. In her laboratory, Mellins led studies of how different genetic variations of MHC II molecules contribute to a vulnerability to autoimmune disease. Her team investigated how immune cells called monocytes function in a variety of immune diseases, including Still disease, rheumatoid arthritis, psoriatic arthritis, asthma and PANS, seeking clues to disease pathogenesis and biomarkers.
Anybody who had gone through Betsy’s lab had a friend and mentor for life.
“Betsy had a relentless work ethic, always excited by new findings and developing the next question she wanted to solve,” said Mark Kay , MD, PhD, professor of pediatrics and of genetics. “She was a remarkable colleague in so many ways. Not only will I miss our scientific discussions but our friendship that included long talks about world events, music, art and our families.”
“She was the sort of person who would call and say, ‘Could we have coffee? I have this interesting idea and want to run it by you,’” said PJ Utz , MD, professor of medicine. “She was motivated by innate curiosity and a desire to discover.”
“Her North Star, within her research, was understanding the mechanism of disease and trying to look differently at the illnesses so you can cure them if possible,” Saper said.
One of Mellins’ most notable research findings, in collaboration with Saper, focused on drugs introduced in the early 2010s to block IL-1 and IL-6, molecules involved in the immune phenomenon known as “cytokine storm.” The drugs are used for Still disease, which is characterized by high fevers, joint inflammation and a risk of cytokine storm.
In 2013, reports emerged of mysterious, life-threatening lung problems in some Still patients, whose lungs showed unique changes in imaging and pathology. Mellins led detective work to demonstrate that children who experienced the problem had specific variants of certain MHC II genes. The discovery has enabled doctors to screen children for the genetic marker and use these drugs with caution in vulnerable children, monitoring them closely for early signs of the complication, or in some cases, avoiding giving the drugs to high-risk patients.
“It really changed the way we practice,” said Christy Sandborg , MD, professor emeritus of pediatrics, who was hired as clinical chief of pediatric rheumatology in 1997 and worked closely with Mellins.
Building a nascent field
In the late 1990s, soon after arriving at Stanford Medicine, Mellins recognized the need for a national research network in pediatric rheumatology and founded the Childhood Arthritis and Rheumatology Research Alliance (CARRA).
“Pediatric rheumatology at that period was very under-resourced, and it was impossible to do research on rare autoimmune diseases,” Sandborg said.
After attending an Arthritis Foundation special workshop on research in juvenile arthritis, Mellins was motivated to build a network that would offer funding and other resources for basic science in pediatric rheumatology and make it easier to conduct large, multicenter clinical trials. She began working with colleagues at Stanford and across the country on the project.
“Everyone in the field was so excited,” Sandborg said.
Among her contributions to the organization, Mellins wrote the first grant to the Arthritis Foundation for CARRA’s support and served as its first chairperson and head of the CARRA research grant committee.
“She recognized that you can do strong clinical work, but if you don’t do lab research, you don’t advance the field,” Sandborg said. “She contributed greatly to that over many years.”
CARRA is now a nonprofit organization with a membership of more than 400 pediatric rheumatologists and researchers across North America. It has garnered millions of dollars in funding and hosts a biorepository that scientists use for rheumatology research.
A devoted mentor
Mellins worked behind the scenes at Stanford Medicine to help recruit incoming medical students who wanted to pursue careers as physician-scientists, which she saw as an essential component of expanding the pipeline of medical researchers, Utz said. “She made a big difference in the recruitment and retention of women in our training programs, including the immunology PhD program and more recently the Berg Scholars Program,” he added.
Frankovich, who was mentored by Mellins early in her career, continued to rely on her advice as they studied PANS, an abrupt-onset, severe psychiatric condition that is thought to be secondary to post-infectious inflammation.
“Her advice was helpful because it wasn’t just about publishing a novel finding; it was about systematically deciphering a complex condition,” Frankovich said. “She was a realist: She knew that it would be very difficult and complex to prove that PANS was secondary to inflammation.”
She cared about the patients I was telling her about, the parents whose kids were affected and the researchers working on the disease.
One big obstacle to studying PANS was access to the parts of the brain involved — the investigators couldn’t biopsy affected brain regions, for example.
“Betsy said, ‘It will be more work to prove an immune hypothesis because we won’t have tissue,’” Frankovich said. “She helped me figure out the biological principles we needed to prove. And then she supported me all the way through each obstacle. When I came to her with a hypothesis, she would say, ‘We can do this, we can solve it,’ and then she would immediately dive into the next steps.
“All along the way she just cared: She cared about the patients I was telling her about, the parents whose kids were affected and the researchers working on the disease.”
Mellins and her husband loved to host visitors from all over the world, their daughter said, including relatives from the United Kingdom and Israel.
“For a recent wedding anniversary, I made them an innkeeper’s book, an album of notes and pictures from the many people who have stayed with them,” Mendelman said. “She deeply valued human connection and she was so gifted at it; she would connect, and stay connected, with anyone.”
Riding his bicycle past Mellins’ and Mendelman’s home on the Stanford campus, Utz could always tell when they had their grown children, grandchildren or other relatives or friends visiting. “There would be five or six cars parked outside,” he said, chuckling.
Mellins and her husband loved live events, attending theater, sports, jazz concerts and modern dance with equal gusto. They didn’t have a TV; instead, “They went to endless performances all over the Bay Area,” Lisa Mendelman said. “Her vision of a balanced life included all of these humanistic things.”
Mellins also loved to read and stayed up to date on contemporary art and literature, deriving great pleasure from finding exactly the right book to give a friend or colleague.
“She drove all the way to my house to give me a copy of the brand-new, hot-off-the-press pediatric rheumatology book,” Frankovich said, adding, “I thought, ‘How did you know this was the book I wanted?’”
Mellins is survived by her husband, Paul Mendelman; son, Jeff Mendelman; daughter, Lisa Mendelman (David Jack); stepson, Adam Mendelson (Catherine Wallis); grandchildren, Oscar and Aila Mendelman Jack; brother Tom Mellins (Judy Weinstein); and nephew, Sam Mellins.
Donations in her honor can be made to a designated fund at the Lucile Packard Foundation for Children’s Health: https://my.supportlpch.org/Mellins .

About Stanford Medicine
Stanford Medicine is an integrated academic health system comprising the Stanford School of Medicine and adult and pediatric health care delivery systems. Together, they harness the full potential of biomedicine through collaborative research, education and clinical care for patients. For more information, please visit med.stanford.edu .
Artificial intelligence
Exploring ways AI is applied to health care

Bank Runs, Fragility, and Regulation
We examine banking regulation in a macroeconomic model of bank runs. We construct a general equilibrium model where banks may default because of fundamental or self-fulfilling runs. With only fundamental defaults, we show that the competitive equilibrium is constrained efficient. However, when banks are vulnerable to runs, banks’ leverage decisions are not ex-ante optimal: individual banks do not internalize that higher leverage makes other banks more vulnerable. The theory calls for introducing minimum capital requirements, even in the absence of bailouts.
The views expressed herein are those of the authors and not necessarily those of the Federal Reserve Bank of Minneapolis, the Federal Reserve System, or the National Bureau of Economic Research.
MARC RIS BibTeΧ
Download Citation Data
More from NBER
In addition to working papers , the NBER disseminates affiliates’ latest findings through a range of free periodicals — the NBER Reporter , the NBER Digest , the Bulletin on Retirement and Disability , the Bulletin on Health , and the Bulletin on Entrepreneurship — as well as online conference reports , video lectures , and interviews .


IMAGES
VIDEO
COMMENTS
122 The Best Genetics Research Topics For Projects. The study of genetics takes place across different levels of the education system in academic facilities all around the world. It is an academic discipline that seeks to explain the mechanism of heredity and genes in living organisms. First discovered back in the 1850s, the study of genetics ...
Learn more about Genetics as a subject, how to choose one of the 119 genetics research topics in this field, and what you need to know while writing. Toll-free: +1 (877) 401-4335. Order Now. About; Prices; ... The next step after choosing genetics research paper topics is to identify relevant sources that will back your research. For Genetics ...
1. Behavioural genetics: Twin and family studies: Measured genetic variants: Quasi-experimental designs: Genetic influences on behaviour: Nature of environmental influence: Nature of genetic influence: Psychiatric genetics: 2. Cytogenetics: Karyotyping: Banding technique: Comparative genome hybridization: FISH (fluorescent in situ hybridization ...
These are controversial topics genetics students can find worth exploring. Nevertheless, prepare to research extensively to compose a winning paper about any of these ideas. Human Genetics Topics For Research Papers. Human genetics entails studying inheritance in human beings. Here are some of the exciting human genetics topics for research papers.
RSS Feed. Genetics is the branch of science concerned with genes, heredity, and variation in living organisms. It seeks to understand the process of trait inheritance from parents to offspring ...
In 1987, the New York Times Magazine characterized the Human Genome Project as the "biggest, costliest, most provocative biomedical research project in history." 2 But in the years between the ...
Combating Antimicrobial Resistant Infectious Diseases in Low- and Middle-Income Countries: Evaluating the Contribution of Genetics and Genomics. The most cited genetics and heredity journal, which advances our understanding of genes from humans to plants and other model organisms. It highlights developments in the function and variability o...
The Journal of Human Genetics is proud to feature "Hot topics in human genetics" - a limited-time web focus bringing together research spanning three highly talked about topics, including ...
Genetics Research Paper Topics. See our list of genetics research paper topics. Genetics is the branch of biology concerned with the science of heredity. The term heredity refers to the way in which specific characteristics are transmitted from one generation to the next. For example, we know that a tall mother and a tall father tend to have ...
Heterozygous CAPZA2 mutations cause global developmental delay, hypotonia with epilepsy: a case report and the literature review. Xiao-Man Zhang. Kai-Li Xu. Shi-Yue Mei. Article 19 Feb 2024.
In the early 1900s, focusing on the evolution of genetic variants in the population, R. A. Fisher, S. Wright, and J. B. S. Haldane made fundamental theoretical contributions to population genetics (Provine 1971), Fisher in his 1922 paper (Fisher 1922), which was the first to introduce diffusion equations into population genetics, and Haldane in ...
Research Topics. The Center for Genetic Medicine's faculty members represent 33 departments or programs across three Northwestern University schools and three Feinberg-affiliated healthcare institutions. Faculty use genetics and molecular genetic approaches to understand biological processes for a diverse range of practical and clinical ...
Genetic Diseases: Sickle Cell Anemia. This genetic disorder research paper aims to elucidate the underlying molecular causes of SCA as well as its symptoms, inheritance, treatment, diagnosis, and prevalence in certain populations. Genetically Identical Twins and Different Disease Risk.
The role of biotechnology in developing non-invasive prenatal genetic testing methods. Genetic engineering for the development of novel enzymes for industrial applications. Investigating the potential of xenotransplantation in addressing organ donor shortages. The use of biotechnology in creating personalised cancer vaccines.
204 Genetics Research Topics & Essay Questions for College and High School. Genetics studies how genes and traits pass from generation to generation. It has practical applications in many areas, such as genetic engineering, gene therapy, gene editing, and genetic testing. If you're looking for exciting genetics topics for presentation, you ...
You can find research titles in areas like cytogenetics, genomics, clinical genetics, and genetics counseling here, as well as appropriate biology topics to write about. These topics will give you solid research ideas and a perfectly formulated paper: Discuss the kinds of genetically transmitted diseases.
Introduction. The past two decades have seen major shifts in our technical ability to sequence genetic information at scale. Historically, genetic testing tended to consist of either highly detailed molecular testing of nominated single genes, or broad genome-wide dosage screening at low resolution, for example karyotyping [1,2].Genome sequencing was too slow and too expensive to be used in ...
As we reflect on these diverse contributions in the Special Issue "Latest Advances in Molecular Genetics and Genomics (2023)", a common thread emerges: the intricate link between genetics, molecular mechanisms, and disease. These review papers collectively underscore the remarkable strides we have taken in unraveling the molecular ...
Genetics research articles from across Nature Portfolio. Genetics research is the scientific discipline concerned with the study of the role of genes in traits such as the development of disease ...
Challenges and Prospects for Conservation Genetics at XXI Century. Conservation genetics is a fundamental discipline for planning actions for the species and ecosystems management, conservation, and sustainable use. Initially, this work was carried out through collaborative efforts of research groups, sometimes from...
Credit: Vegetable Research (2024). DOI: 10.48130/vegres-0024-0005. A research team has elucidated the role of polyploidy in the evolution and breeding of vegetable crops, leveraging advanced ...
They studied genetic samples from 234 patients who had been recruited for research over an 18-year span. The patients had exotropia, while their family members either had or didn't have strabismus.
Overview of the cohorts, analysis pipeline and genetic correlations among the sites. Credit: Nature Human Behaviour (2024). DOI: 10.1038/s41562-024-01851-6
Research in genetics and infectious diseases (ID) presents novel configurations of ethical, legal, and social issues (ELSIs) related to the intersection of genetics with public health regulations and the control of transmissible diseases. ... This paper is submitted on behalf of the Working Group on ELSI, Biobanks, Infectious Diseases, and ...
The Santa Barbara Sense of Direction Scale is widely used in navigation research. Studies suggest that people are fairly accurate at evaluating their own sense of direction. Personality, too, appears to play a role in developing navigational ability. "To get good at navigating, you have to be willing to explore," says Uttal.
Genomics is the study of the full genetic complement of an organism (the genome). It employs recombinant DNA, DNA sequencing methods, and bioinformatics to sequence, assemble, and analyse the ...
Thanks to a genetic breakthrough, a rare rhino species may be rescued from extinction New research from the San Diego Zoo using just 12 frozen cells suggests we can protect the northern white rhino
Visualization with AI. Meeting Time: 04:00 PM‑05:15 PM TTh. Instructor: Qianwen Wang. Course Description: This course aims to investigate how visualization techniques and AI technologies work together to enhance understanding, insights, or outcomes. This is a seminar style course consisting of lectures, paper presentation, and interactive ...
April 12, 2024 - By Erin Digitale. Betsy Mellins "wanted other people to care about the science as much as she cared, because it was science that would lead to better treatments for children," Jennifer Frankovich said. Elizabeth "Betsy" Mellins, MD, professor of pediatrics at the Stanford School of Medicine, died March 24. She was 72.
Working Paper 32341. DOI 10.3386/w32341. Issue Date April 2024. We examine banking regulation in a macroeconomic model of bank runs. We construct a general equilibrium model where banks may default because of fundamental or self-fulfilling runs. With only fundamental defaults, we show that the competitive equilibrium is constrained efficient.